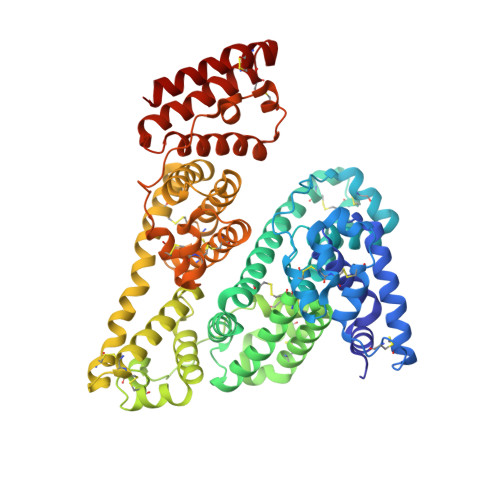
Structural Characterization of a Gold/Serum Albumin Complex.
Pratesi, A., Cirri, D., Fregona, D., Ferraro, G., Giorgio, A., Merlino, A., Messori, L.(2019) Inorg Chem 58: 10616-10619
- PubMed: 31361466 Search on PubMed
- DOI: https://doi.org/10.1021/acs.inorgchem.9b01900
- Primary Citation Related Structures:
6RJV - PubMed Abstract:
The medicinal gold(III) dithiocarbamato complex AuL 12 forms a stable adduct with bovine serum albumin. The crystal structure reveals that a single gold(I) center is bound to Cys34, with the dithiocarbamato ligand being released. To the best of our knowledge, this is the first structure for a gold adduct of serum albumin.
- Laboratory of Metals in Medicine (MetMed), Department of Chemistry "U. Schiff" , University of Florence , via della Lastruccia 3 , 50019 Sesto Fiorentino , Italy.
Organizational Affiliation: