N-locking stabilization of covalent helical peptides: Application to Bfl-1 antagonists.
Baggio, C., Udompholkul, P., Gambini, L., Jossart, J., Salem, A.F., Hakansson, M., Perry, J.J.P., Pellecchia, M.(2020) Chem Biol Drug Des 95: 412-426
- PubMed: 31898401 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/cbdd.13661
- Primary Citation Related Structures:
6RJP - PubMed Abstract:
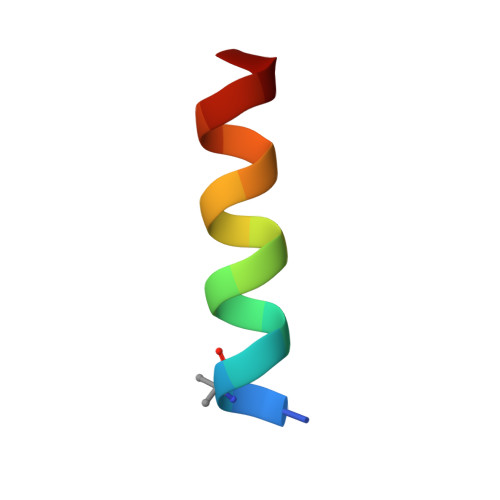
Recently, it was reported that tetrapeptides cyclized via lactam bond between the amino terminus and a glutamic residue in position 4 (termed here N-lock) can nucleate helix formation in longer peptides. We applied such strategy to derive N-locked covalent BH3 peptides that were designed to selectively target the anti-apoptotic protein Bfl-1. The resulting agents were soluble in aqueous buffer and displayed a remarkable (low nanomolar) affinity for Bfl-1 and cellular activity. The crystal structure of the complex between such N-locked covalent peptide and Bfl-1 provided insights on the geometry of the N-locking strategy and of the covalent bond between the agent and Bfl-1.
- Division of Biomedical Sciences, School of Medicine, University of California Riverside, Riverside, CA, USA.
Organizational Affiliation: