Microfluidic protein isolation and sample preparation for high-resolution cryo-EM.
Schmidli, C., Albiez, S., Rima, L., Righetto, R., Mohammed, I., Oliva, P., Kovacik, L., Stahlberg, H., Braun, T.(2019) Proc Natl Acad Sci U S A 116: 15007-15012
- PubMed: 31292253 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1907214116
- Primary Citation Related Structures:
6R70, 6R7M - PubMed Abstract:
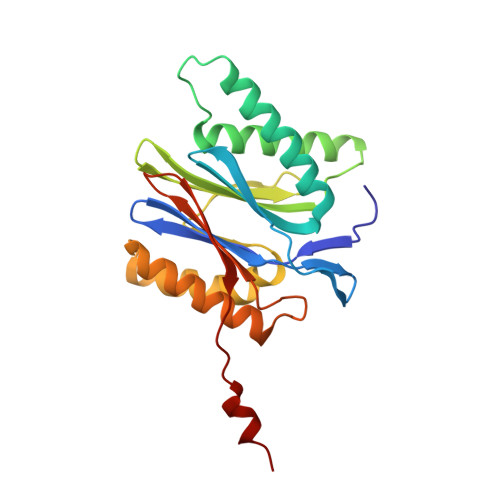
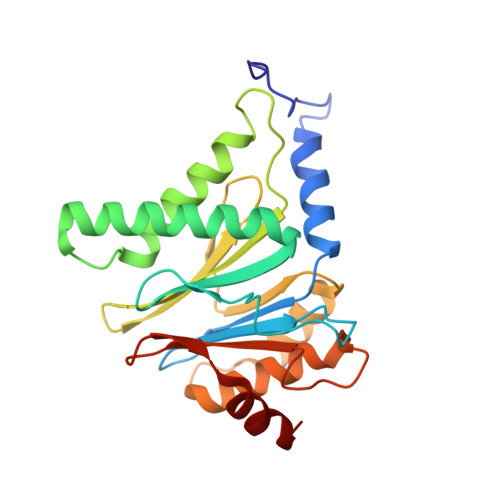
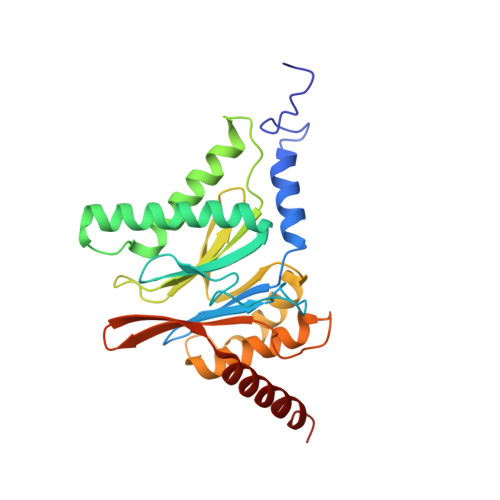
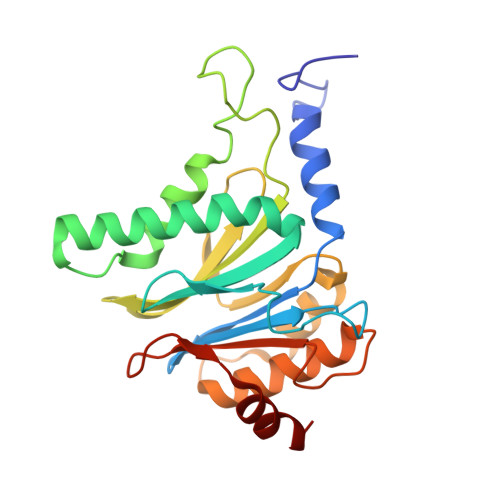
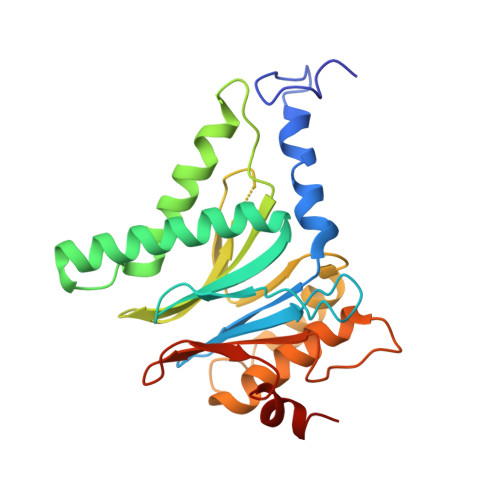
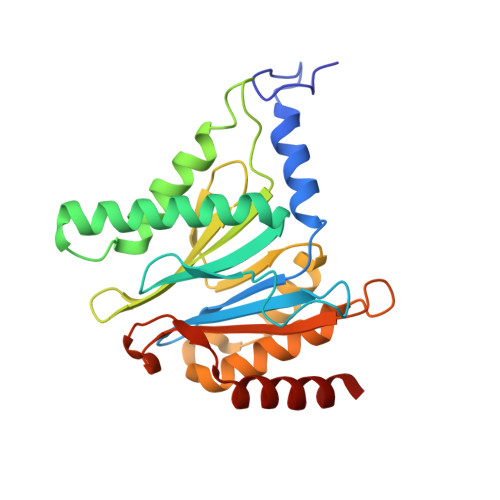
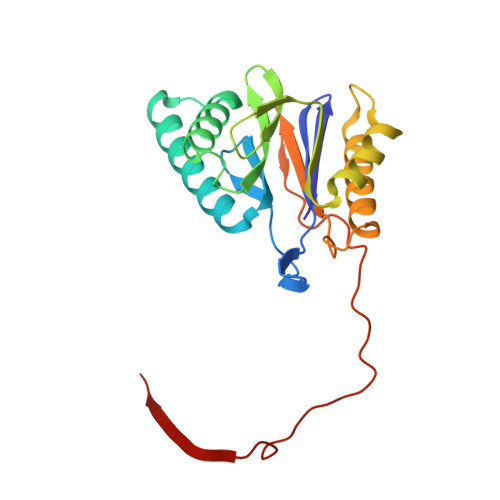
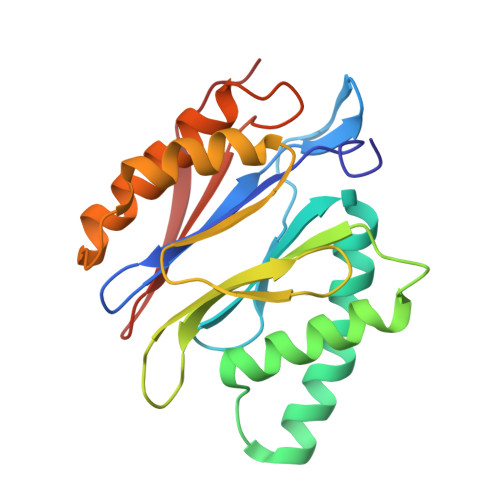
High-resolution structural information is essential to understand protein function. Protein-structure determination needs a considerable amount of protein, which can be challenging to produce, often involving harsh and lengthy procedures. In contrast, the several thousand to a few million protein particles required for structure determination by cryogenic electron microscopy (cryo-EM) can be provided by miniaturized systems. Here, we present a microfluidic method for the rapid isolation of a target protein and its direct preparation for cryo-EM. Less than 1 μL of cell lysate is required as starting material to solve the atomic structure of the untagged, endogenous human 20S proteasome. Our work paves the way for high-throughput structure determination of proteins from minimal amounts of cell lysate and opens more opportunities for the isolation of sensitive, endogenous protein complexes.
- Center for Cellular Imaging and Nanoanalytics, Biozentrum, 4058 Basel, Switzerland.
Organizational Affiliation: