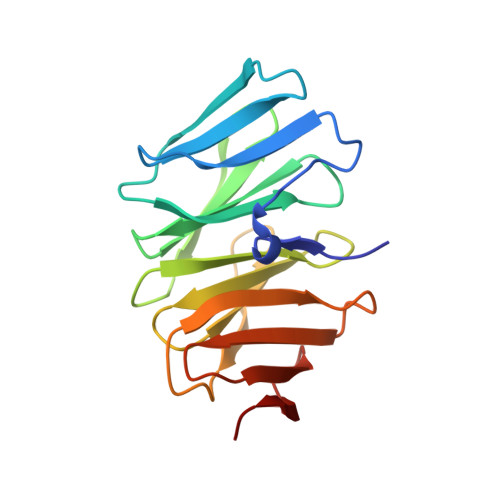
Structural diversity of oligomeric beta-propellers with different numbers of identical blades.
Afanasieva, E., Chaudhuri, I., Martin, J., Hertle, E., Ursinus, A., Alva, V., Hartmann, M.D., Lupas, A.N.(2019) Elife 8
- PubMed: 31613220 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.49853
- Primary Citation Related Structures:
6R5X, 6R5Y, 6R5Z, 6R60 - PubMed Abstract:
β-Propellers arise through the amplification of a supersecondary structure element called a blade. This process produces toroids of between four and twelve repeats, which are almost always arranged sequentially in a single polypeptide chain. We found that new propellers evolve continuously by amplification from single blades. We therefore investigated whether such nascent propellers can fold as homo-oligomers before they have been fully amplified within a single chain. One- to six-bladed building blocks derived from two seven-bladed WD40 propellers yielded stable homo-oligomers with six to nine blades, depending on the size of the building block. High-resolution structures for tetramers of two blades, trimers of three blades, and dimers of four and five blades, respectively, show structurally diverse propellers and include a novel fold, highlighting the inherent flexibility of the WD40 blade. Our data support the hypothesis that subdomain-sized fragments can provide structural versatility in the evolution of new proteins.
- Department of Protein Evolution, Max Planck Institute for Developmental Biology, Tübingen, Germany.
Organizational Affiliation: