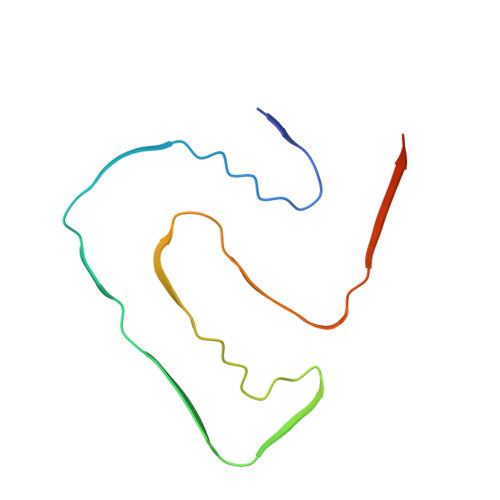
Atomic structure of PI3-kinase SH3 amyloid fibrils by cryo-electron microscopy.
Roder, C., Vettore, N., Mangels, L.N., Gremer, L., Ravelli, R.B.G., Willbold, D., Hoyer, W., Buell, A.K., Schroder, G.F.(2019) Nat Commun 10: 3754-3754
- PubMed: 31434882 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-11320-8
- Primary Citation Related Structures:
6R4R - PubMed Abstract:
High resolution structural information on amyloid fibrils is crucial for the understanding of their formation mechanisms and for the rational design of amyloid inhibitors in the context of protein misfolding diseases. The Src-homology 3 domain of phosphatidyl-inositol-3-kinase (PI3K-SH3) is a model amyloid system that plays a pivotal role in our basic understanding of protein misfolding and aggregation. Here, we present the atomic model of the PI3K-SH3 amyloid fibril with a resolution determined to 3.4 Å by cryo-electron microscopy (cryo-EM). The fibril is composed of two intertwined protofilaments that create an interface spanning 13 residues from each monomer. The model comprises residues 1-77 out of 86 amino acids in total, with the missing residues located in the highly flexible C-terminus. The fibril structure allows us to rationalise the effects of chemically conservative point mutations as well as of the previously reported sequence perturbations on PI3K-SH3 fibril formation and growth.
- Institute of Complex Systems, Structural Biochemistry (ICS-6) and JuStruct, Jülich Center for Structural Biology, Forschungszentrum Jülich, 52425, Jülich, Germany.
Organizational Affiliation: