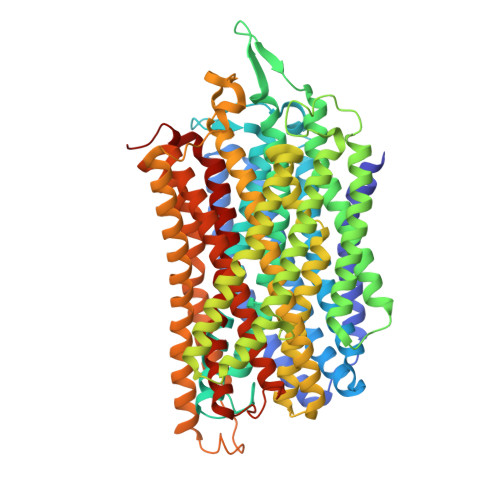
Asymmetry in catalysis byThermotoga maritimamembrane-bound pyrophosphatase demonstrated by a nonphosphorus allosteric inhibitor.
Vidilaseris, K., Kiriazis, A., Turku, A., Khattab, A., Johansson, N.G., Leino, T.O., Kiuru, P.S., Boije Af Gennas, G., Meri, S., Yli-Kauhaluoma, J., Xhaard, H., Goldman, A.(2019) Sci Adv 5: eaav7574-eaav7574
- PubMed: 31131322 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.aav7574
- Primary Citation Related Structures:
6QXA - PubMed Abstract:
Membrane-bound pyrophosphatases are homodimeric integral membrane proteins that hydrolyze pyrophosphate into orthophosphates, coupled to the active transport of protons or sodium ions across membranes. They are important in the life cycle of bacteria, archaea, plants, and parasitic protists, but no homologous proteins exist in vertebrates, making them a promising drug target. Here, we report the first nonphosphorus allosteric inhibitor of the thermophilic bacterium Thermotoga maritima membrane-bound pyrophosphatase and its bound structure together with the substrate analog imidodiphosphate. The unit cell contains two protein homodimers, each binding a single inhibitor dimer near the exit channel, creating a hydrophobic clamp that inhibits the movement of β-strand 1-2 during pumping, and thus prevents the hydrophobic gate from opening. This asymmetry of inhibitor binding with respect to each homodimer provides the first clear structural demonstration of asymmetry in the catalytic cycle of membrane-bound pyrophosphatases.
- Research Program in Molecular and Integrative Biosciences, University of Helsinki, Helsinki, Finland.
Organizational Affiliation: