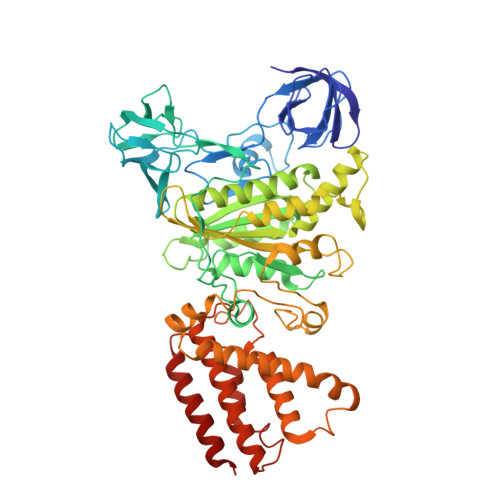
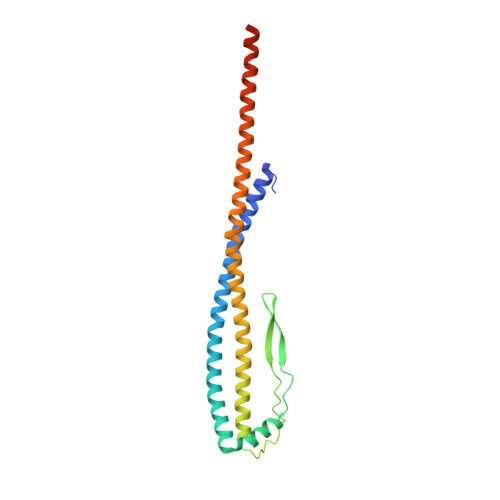
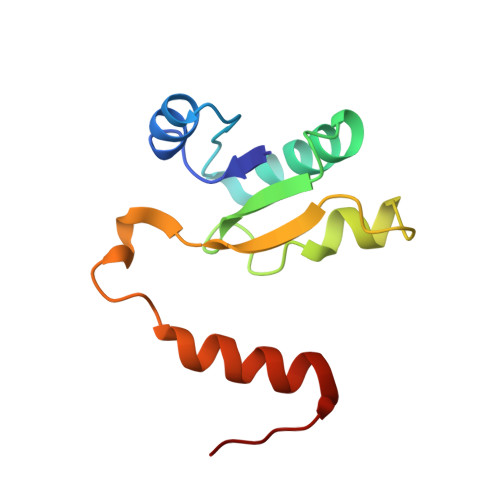
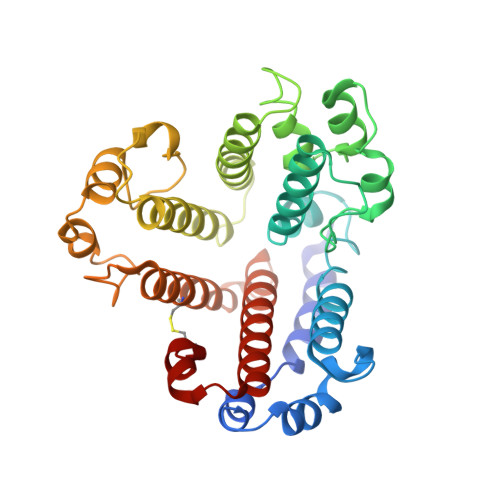
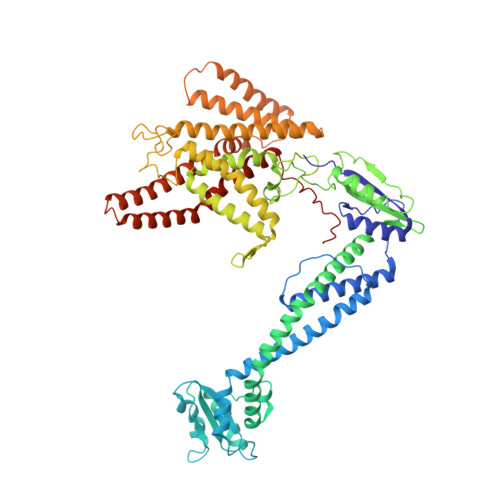
Structure and conformational plasticity of the intact Thermus thermophilus V/A-type ATPase.
Zhou, L., Sazanov, L.A.(2019) Science 365
- PubMed: 31439765 Search on PubMed
- DOI: https://doi.org/10.1126/science.aaw9144
- Primary Citation Related Structures:
6QUM, 6R0W, 6R0Y, 6R0Z, 6R10 - PubMed Abstract:
V (vacuolar)/A (archaeal)-type adenosine triphosphatases (ATPases), found in archaea and eubacteria, couple ATP hydrolysis or synthesis to proton translocation across the plasma membrane using the rotary-catalysis mechanism. They belong to the V-type ATPase family, which differs from the mitochondrial/chloroplast F-type ATP synthases in overall architecture. We solved cryo-electron microscopy structures of the intact Thermus thermophilus V/A-ATPase, reconstituted into lipid nanodiscs, in three rotational states and two substates. These structures indicate substantial flexibility between V 1 and V o in a working enzyme, which results from mechanical competition between central shaft rotation and resistance from the peripheral stalks. We also describe details of adenosine diphosphate inhibition release, V 1 -V o torque transmission, and proton translocation, which are relevant for the entire V-type ATPase family.
- Institute of Science and Technology Austria, Klosterneuberg 3400, Austria.
Organizational Affiliation: