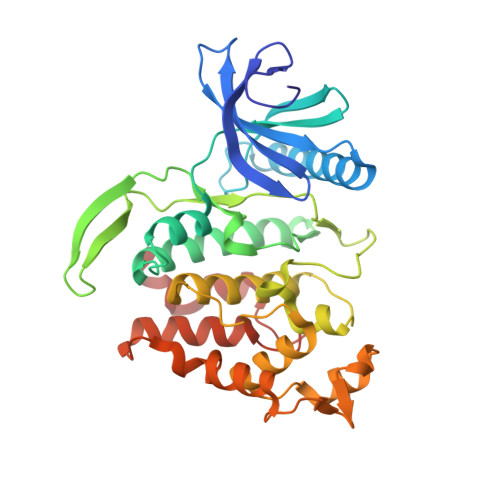
Expression, purification and crystallization of CLK1 kinase - A potential target for antiviral therapy.
Dekel, N., Eisenberg-Domovich, Y., Karlas, A., Meyer, T.F., Bracher, F., Lebendiker, M., Danieli, T., Livnah, O.(2020) Protein Expr Purif 176: 105742-105742
- PubMed: 32866611 Search on PubMed
- DOI: https://doi.org/10.1016/j.pep.2020.105742
- Primary Citation Related Structures:
6QTY - PubMed Abstract:
Cdc-like kinase 1 (CLK1) is a dual-specificity kinase capable of autophosphorylation on tyrosine residues and Ser/Thr phosphorylation of its substrates. CLK1 belongs to the CLK kinase family that regulates alternative splicing through phosphorylation of serine-arginine rich (SR) proteins. Recent studies have demonstrated that CLK1 has an important role in the replication of influenza A and chikungunya viruses. Furthermore, CLK1 was found to be relevant for the replication of HIV-1 and the West Nile virus, making CLK1 an interesting cellular candidate for the development of a host-directed antiviral therapy that might be efficient for treatment of newly emerging viruses. We describe here our attempts and detailed procedures to obtain the recombinant kinase domain of CLK1 in suitable amounts for crystallization in complex with specific inhibitors. The key solution for the reproducibility of crystals resides in devising and refining expression and purification protocols leading to homogeneous protein. Co-expression of CLK1 with λ-phosphatase and careful purification has yielded crystals of CLK1 complexed with the KH-CB19 inhibitor that diffracted to 1.65 Å. These results paved the path to the screening of more structures of CLK1 complexed compounds, leading to further optimization of their inhibitory activity. Moreover, since kinases are desired targets in numerous pathologies, the approach we report here, the co-expression of kinases with λ-phosphatase, previously used in other kinases, can be adopted as a general protocol in numerous kinase targets for obtaining reproducible and homogenic non-phosphorylated (inactive) forms suitable for biochemical and structural studies thus facilitating the development of novel inhibitors.
- The Wolfson Centre for Applied Structural Biology, The Hebrew University of Jerusalem, Jerusalem 91904, Israel.
Organizational Affiliation: