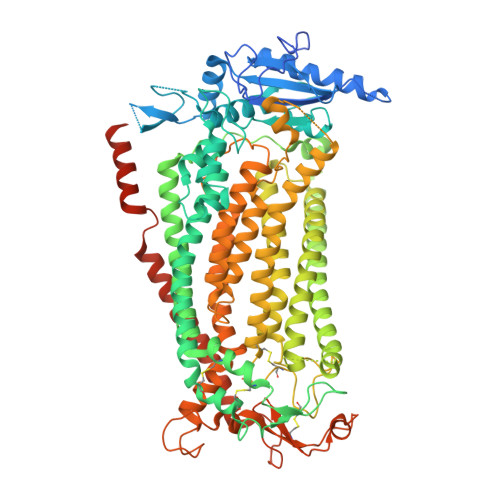
Cryo-EM structures and functional characterization of the murine lipid scramblase TMEM16F.
Alvadia, C., Lim, N.K., Clerico Mosina, V., Oostergetel, G.T., Dutzler, R., Paulino, C.(2019) Elife 8
- PubMed: 30785399 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.44365
- Primary Citation Related Structures:
6QP6, 6QPB, 6QPC, 6QPI - PubMed Abstract:
The lipid scramblase TMEM16F initiates blood coagulation by catalyzing the exposure of phosphatidylserine in platelets. The protein is part of a family of membrane proteins, which encompasses calcium-activated channels for ions and lipids. Here, we reveal features of murine TMEM16F (mTMEM16F) that underlie its function as a lipid scramblase and an ion channel. The cryo-EM data of mTMEM16F in absence and presence of Ca 2+ define the ligand-free closed conformation of the protein and the structure of a Ca 2+ -bound intermediate. Both conformations resemble their counterparts of the scrambling-incompetent anion channel mTMEM16A, yet with distinct differences in the region of ion and lipid permeation. In conjunction with functional data, we demonstrate the relationship between ion conduction and lipid scrambling. Although activated by a common mechanism, both functions appear to be mediated by alternate protein conformations that are at equilibrium in the ligand-bound state.
- Department of Biochemistry, University of Zurich, Zurich, Switzerland.
Organizational Affiliation: