The structural basis for cohesin-CTCF-anchored loops.
Li, Y., Haarhuis, J.H.I., Sedeno Cacciatore, A., Oldenkamp, R., van Ruiten, M.S., Willems, L., Teunissen, H., Muir, K.W., de Wit, E., Rowland, B.D., Panne, D.(2020) Nature 578: 472-476
- PubMed: 31905366
- DOI: https://doi.org/10.1038/s41586-019-1910-z
- Primary Citation Related Structures:
6QNX - PubMed Abstract:
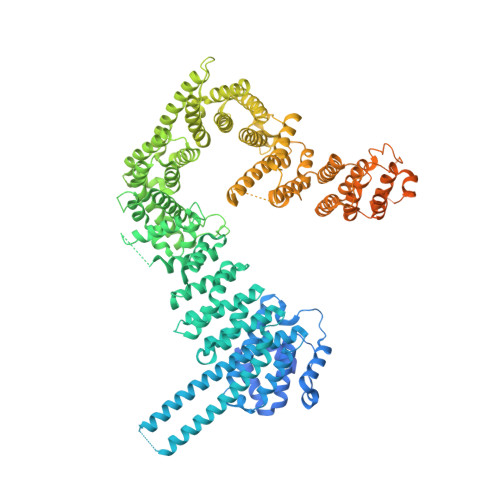
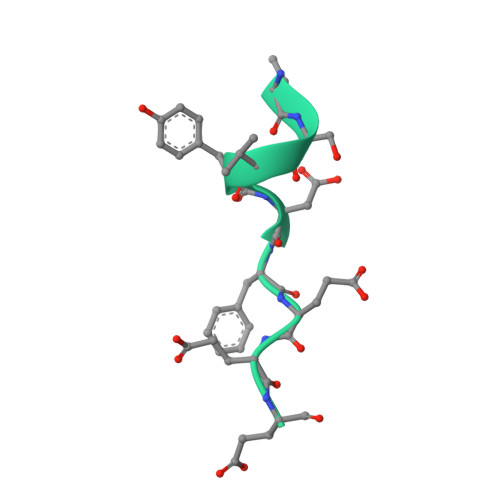
Cohesin catalyses the folding of the genome into loops that are anchored by CTCF 1 . The molecular mechanism of how cohesin and CTCF structure the 3D genome has remained unclear. Here we show that a segment within the CTCF N terminus interacts with the SA2-SCC1 subunits of human cohesin. We report a crystal structure of SA2-SCC1 in complex with CTCF at a resolution of 2.7 Å, which reveals the molecular basis of the interaction. We demonstrate that this interaction is specifically required for CTCF-anchored loops and contributes to the positioning of cohesin at CTCF binding sites. A similar motif is present in a number of established and newly identified cohesin ligands, including the cohesin release factor WAPL 2,3 . Our data suggest that CTCF enables the formation of chromatin loops by protecting cohesin against loop release. These results provide fundamental insights into the molecular mechanism that enables the dynamic regulation of chromatin folding by cohesin and CTCF.
- European Molecular Biology Laboratory, Grenoble, France.
Organizational Affiliation: