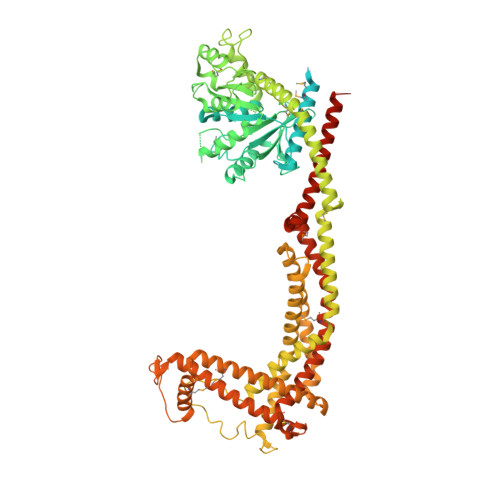
Structure and assembly of the mitochondrial membrane remodelling GTPase Mgm1.
Faelber, K., Dietrich, L., Noel, J.K., Wollweber, F., Pfitzner, A.K., Muhleip, A., Sanchez, R., Kudryashev, M., Chiaruttini, N., Lilie, H., Schlegel, J., Rosenbaum, E., Hessenberger, M., Matthaeus, C., Kunz, S., von der Malsburg, A., Noe, F., Roux, A., van der Laan, M., Kuhlbrandt, W., Daumke, O.(2019) Nature 571: 429-433
- PubMed: 31292547
- DOI: https://doi.org/10.1038/s41586-019-1372-3
- Primary Citation of Related Structures:
6QL4, 6RZT, 6RZU, 6RZV, 6RZW - PubMed Abstract:
Balanced fusion and fission are key for the proper function and physiology of mitochondria 1,2 . Remodelling of the mitochondrial inner membrane is mediated by the dynamin-like protein mitochondrial genome maintenance 1 (Mgm1) in fungi or the related protein optic atrophy 1 (OPA1) in animals 3-5 . Mgm1 is required for the preservation of mitochondrial DNA in yeast 6 , whereas mutations in the OPA1 gene in humans are a common cause of autosomal dominant optic atrophy-a genetic disorder that affects the optic nerve 7,8 . Mgm1 and OPA1 are present in mitochondria as a membrane-integral long form and a short form that is soluble in the intermembrane space. Yeast strains that express temperature-sensitive mutants of Mgm1 9,10 or mammalian cells that lack OPA1 display fragmented mitochondria 11,12 , which suggests that Mgm1 and OPA1 have an important role in inner-membrane fusion. Consistently, only the mitochondrial outer membrane-not the inner membrane-fuses in the absence of functional Mgm1 13 . Mgm1 and OPA1 have also been shown to maintain proper cristae architecture 10,14 ; for example, OPA1 prevents the release of pro-apoptotic factors by tightening crista junctions 15 . Finally, the short form of OPA1 localizes to mitochondrial constriction sites, where it presumably promotes mitochondrial fission 16 . How Mgm1 and OPA1 perform their diverse functions in membrane fusion, scission and cristae organization is at present unknown. Here we present crystal and electron cryo-tomography structures of Mgm1 from Chaetomium thermophilum. Mgm1 consists of a GTPase (G) domain, a bundle signalling element domain, a stalk, and a paddle domain that contains a membrane-binding site. Biochemical and cell-based experiments demonstrate that the Mgm1 stalk mediates the assembly of bent tetramers into helical filaments. Electron cryo-tomography studies of Mgm1-decorated lipid tubes and fluorescence microscopy experiments on reconstituted membrane tubes indicate how the tetramers assemble on positively or negatively curved membranes. Our findings convey how Mgm1 and OPA1 filaments dynamically remodel the mitochondrial inner membrane.
- Crystallography, Max-Delbrück-Centrum for Molecular Medicine, Berlin, Germany. katja.faelber@mdc-berlin.de.
Organizational Affiliation: