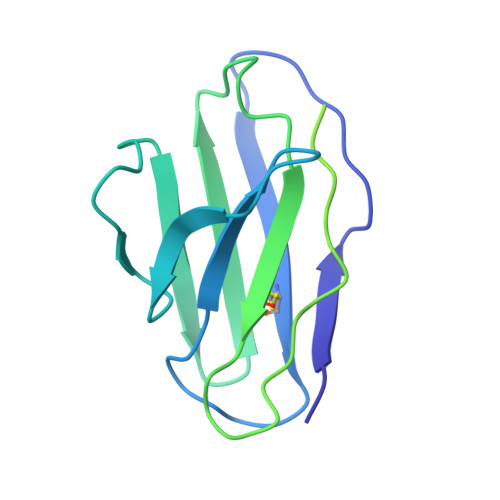
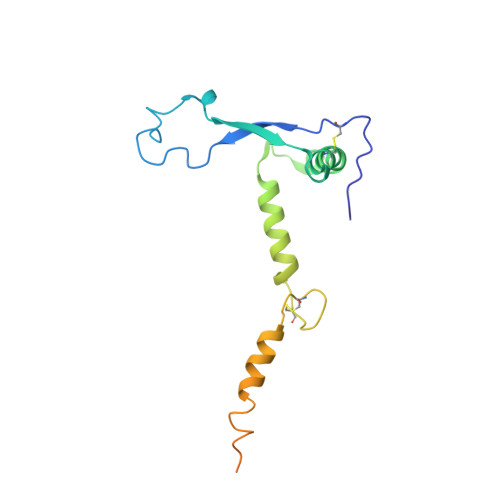
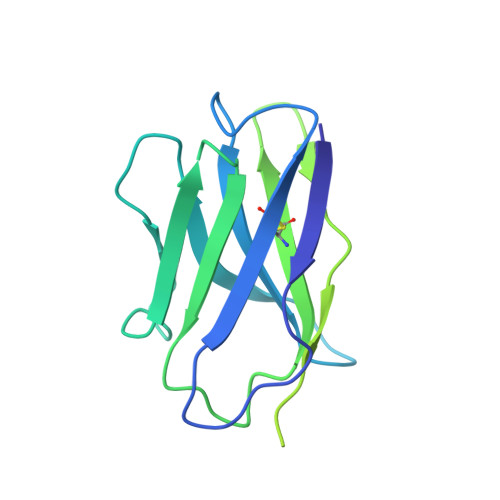
rVSV-ZEBOV induces a polyclonal and convergent B cell response with potent Ebola virus-neutralizing antibodies
Diskin, R., Cohen-Dvashi, H., Ehrhardt, S., Zehner, M., Krahling, V., Kreer, C., Dahlke, C., Ercanoglu, M.S., Gruell, H., Addo, M.M., Becker, S., Klein, F.(2019) Nat Med