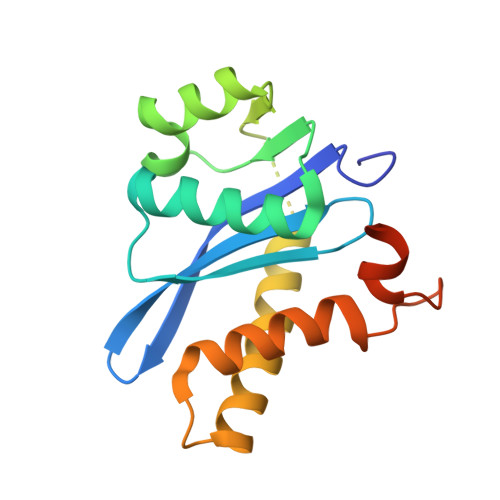
Cryo-EM structure of the deltaretroviral intasome in complex with the PP2A regulatory subunit B56 gamma.
Barski, M.S., Minnell, J.J., Hodakova, Z., Pye, V.E., Nans, A., Cherepanov, P., Maertens, G.N.(2020) Nat Commun 11: 5043-5043
- PubMed: 33028863 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-18874-y
- Primary Citation Related Structures:
6QBT, 6QBV, 6QBW, 6TJU, 6TOQ, 7PEL - PubMed Abstract:
Human T-cell lymphotropic virus type 1 (HTLV-1) is a deltaretrovirus and the most oncogenic pathogen. Many of the ~20 million HTLV-1 infected people will develop severe leukaemia or an ALS-like motor disease, unless a therapy becomes available. A key step in the establishment of infection is the integration of viral genetic material into the host genome, catalysed by the retroviral integrase (IN) enzyme. Here, we use X-ray crystallography and single-particle cryo-electron microscopy to determine the structure of the functional deltaretroviral IN assembled on viral DNA ends and bound to the B56γ subunit of its human host factor, protein phosphatase 2 A. The structure reveals a tetrameric IN assembly bound to two molecules of the phosphatase via a conserved short linear motif. Insight into the deltaretroviral intasome and its interaction with the host will be crucial for understanding the pattern of integration events in infected individuals and therefore bears important clinical implications.
- Imperial College London, St Mary's Hospital, Department of Infectious Disease, Section of Virology, Norfolk Place, London, W2 1PG, UK.
Organizational Affiliation: