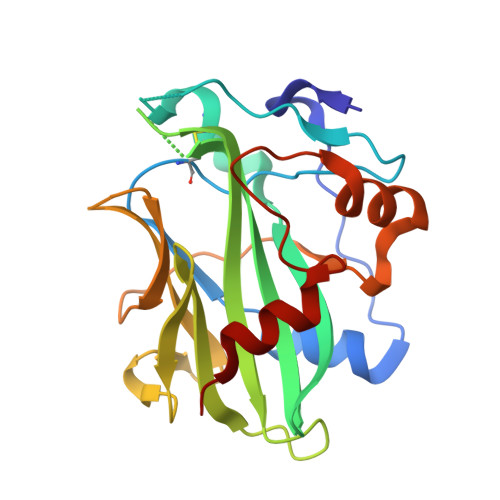
Molecular basis for functional diversity among microbial Nep1-like proteins.
Lenarcic, T., Pirc, K., Hodnik, V., Albert, I., Borisek, J., Magistrato, A., Nurnberger, T., Podobnik, M., Anderluh, G.(2019) PLoS Pathog 15: e1007951-e1007951
- PubMed: 31479498 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1007951
- Primary Citation Related Structures:
6QBD, 6QBE - PubMed Abstract:
Necrosis and ethylene-inducing peptide 1 (Nep1)-like proteins (NLPs) are secreted by several phytopathogenic microorganisms. They trigger necrosis in various eudicot plants upon binding to plant sphingolipid glycosylinositol phosphorylceramides (GIPC). Interestingly, HaNLP3 from the obligate biotroph oomycete Hyaloperonospora arabidopsidis does not induce necrosis. We determined the crystal structure of HaNLP3 and showed that it adopts the NLP fold. However, the conformations of the loops surrounding the GIPC headgroup-binding cavity differ from those of cytotoxic Pythium aphanidermatum NLPPya. Essential dynamics extracted from μs-long molecular dynamics (MD) simulations reveals a limited conformational plasticity of the GIPC-binding cavity in HaNLP3 relative to toxic NLPs. This likely precludes HaNLP3 binding to GIPCs, which is the underlying reason for the lack of toxicity. This study reveals that mutations at key protein regions cause a switch between non-toxic and toxic phenotypes within the same protein scaffold. Altogether, these data provide evidence that protein flexibility is a distinguishing trait of toxic NLPs and highlight structural determinants for a potential functional diversification of non-toxic NLPs utilized by biotrophic plant pathogens.
- Department of Molecular Biology and Nanobiotechnology, National Institute of Chemistry, Hajdrihova, Ljubljana, Slovenia.
Organizational Affiliation: