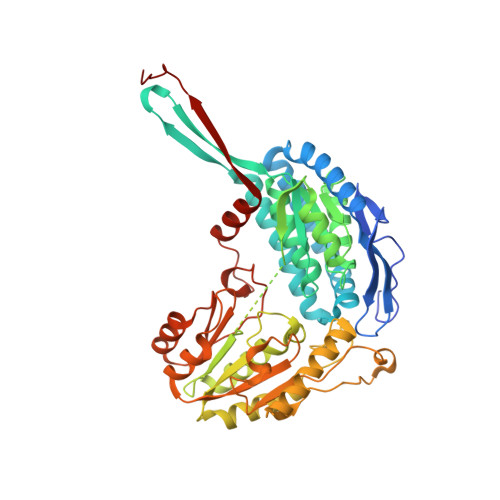
Kinetic and structural analysis of human ALDH9A1.
Koncitikova, R., Vigouroux, A., Kopecna, M., Sebela, M., Morera, S., Kopecny, D.(2019) Biosci Rep 39
- PubMed: 30914451
- DOI: https://doi.org/10.1042/BSR20190558
- Primary Citation of Related Structures:
6QAK, 6QAO, 6QAP - PubMed Abstract:
Aldehyde dehydrogenases (ALDHs) constitute a superfamily of NAD(P) + -dependent enzymes, which detoxify aldehydes produced in various metabolic pathways to the corresponding carboxylic acids. Among the 19 human ALDHs, the cytosolic ALDH9A1 has so far never been fully enzymatically characterized and its structure is still unknown. Here, we report complete molecular and kinetic properties of human ALDH9A1 as well as three crystal forms at 2.3, 2.9, and 2.5 Å resolution. We show that ALDH9A1 exhibits wide substrate specificity to aminoaldehydes, aliphatic and aromatic aldehydes with a clear preference for γ -trimethylaminobutyraldehyde (TMABAL). The structure of ALDH9A1 reveals that the enzyme assembles as a tetramer. Each ALDH monomer displays a typical ALDHs fold composed of an oligomerization domain, a coenzyme domain, a catalytic domain, and an inter-domain linker highly conserved in amino-acid sequence and folding. Nonetheless, structural comparison reveals a position and a fold of the inter-domain linker of ALDH9A1 never observed in any other ALDH so far. This unique difference is not compatible with the presence of a bound substrate and a large conformational rearrangement of the linker up to 30 Å has to occur to allow the access of the substrate channel. Moreover, the αβE region consisting of an α-helix and a β-strand of the coenzyme domain at the dimer interface are disordered, likely due to the loss of interactions with the inter-domain linker, which leads to incomplete β-nicotinamide adenine dinucleotide (NAD + ) binding pocket.
- Department of Protein Biochemistry and Proteomics, Centre of the Region Haná for Biotechnological and Agricultural Research, Faculty of Science, Palacký University, Šlechtitelů 27, CZ-783 71 Olomouc, Czech Republic.
Organizational Affiliation: