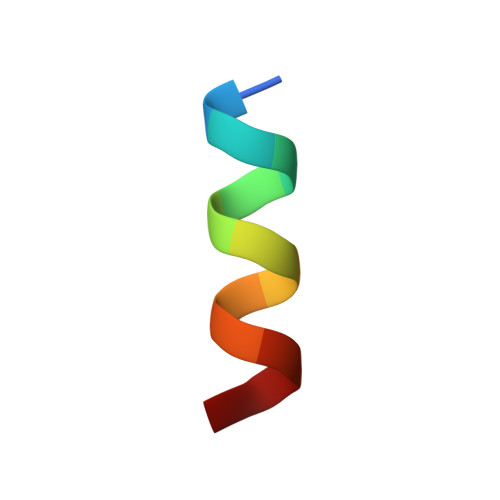
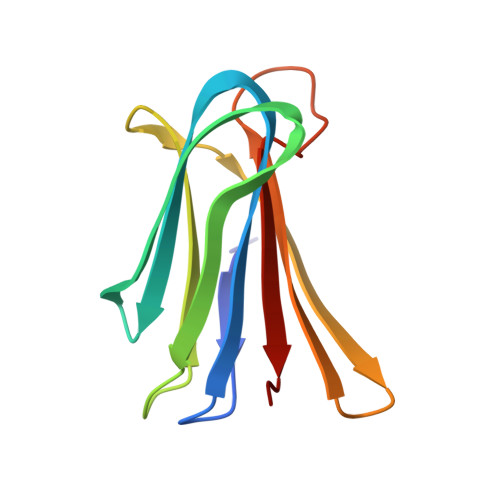
X-ray Crystal Structures of Short Antimicrobial Peptides as Pseudomonas aeruginosa Lectin B Complexes.
Baeriswyl, S., Gan, B.H., Siriwardena, T.N., Visini, R., Robadey, M., Javor, S., Stocker, A., Darbre, T., Reymond, J.L.(2019) ACS Chem Biol 14: 758-766
- PubMed: 30830745 Search on PubMed
- DOI: https://doi.org/10.1021/acschembio.9b00047
- Primary Citation Related Structures:
6Q6W, 6Q6X, 6Q77, 6Q79, 6Q85, 6Q86, 6Q87, 6Q8D, 6Q8G, 6Q8H, 6S5P - PubMed Abstract:
Herein, we report X-ray crystal structures of 11-13 residue antimicrobial peptides (AMPs) active against Pseudomonas aeruginosa as complexes of fucosylated d-enantiomeric sequences with the P. aeruginosa lectin LecB. These represent the first crystal structures of short AMPs. In 24 individual structures of eight different peptides, we found mostly α-helices assembled as two-helix or four-helix bundles with a hydrophobic core and cationic residues pointing outside. Two of the analogs formed an extended structure engaging in multiple contacts with the lectin. Molecular dynamics (MD) simulations showed that α-helices are stabilized by bundle formation and suggested that the N-terminal acyl group present in the linker to the fucosyl group can extend the helix by one additional H-bond and increase α-helix amphiphilicity. Investigating N-terminal acylation led to AMPs with equivalent and partly stronger antibacterial effects compared to the free peptide.
- Department of Chemistry and Biochemistry , University of Bern , Freiestrasse 3 , 3012 Bern , Switzerland.
Organizational Affiliation: