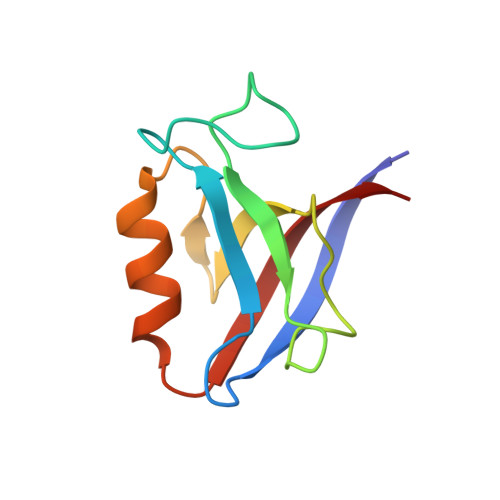
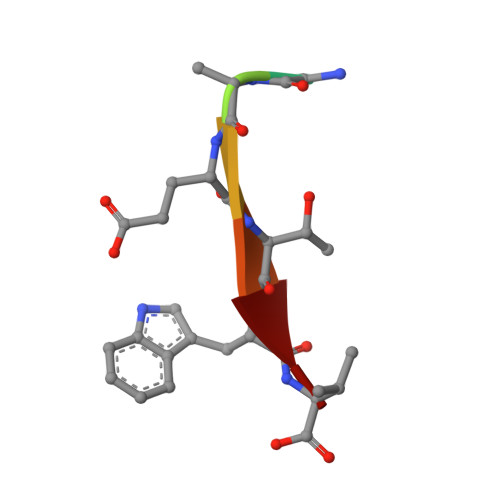
Comprehensive analysis of all evolutionary paths between two divergent PDZ domain specificities.
Teyra, J., Ernst, A., Singer, A., Sicheri, F., Sidhu, S.S.(2020) Protein Sci 29: 433-442
- PubMed: 31654425 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.3759
- Primary Citation Related Structures:
6Q0M, 6Q0N, 6Q0U - PubMed Abstract:
To understand the molecular evolution of functional diversity in protein families, we comprehensively investigated the consequences of all possible mutation combinations separating two peptide-binding domains with highly divergent specificities. We analyzed the Erbin PDZ domain (Erbin-PDZ), which exhibits canonical type I specificity, and a synthetic Erbin-PDZ variant (E-14) that differs at six positions and exhibits an atypical specificity that closely resembles that of the natural Pdlim4 PDZ domain (Pdlim4-PDZ). We constructed a panel of 64 PDZ domains covering all possible transitions between Erbin-PDZ and E-14 (i.e., the panel contained variants with all possible combinations of either the Erbin-PDZ or E-14 sequence at the six differing positions). We assessed the specificity profiles of the 64 PDZ domains using a C-terminal phage-displayed peptide library containing all possible genetically encoded heptapeptides. The specificity profiles clustered into six distinct groups, showing that intermediate domains can be nodes for the evolution of divergent functions. Remarkably, three substitutions were sufficient to convert the specificity of Erbin-PDZ to that of Pdlim4-PDZ, whereas Pdlim4-PDZ contains 71 differences relative to Erbin-PDZ. X-ray crystallography revealed the structural basis for specificity transition: a single substitution in the center of the binding site, supported by contributions from auxiliary substitutions, altered the main chain conformation of the peptide ligand to resemble that of ligands bound to Pdlim4-PDZ. Our results show that a very small set of mutations can dramatically alter protein specificity, and these findings support the hypothesis whereby complex protein functions evolve by gene duplication followed by cumulative mutations.
- The Donnelly Centre, University of Toronto, Toronto, Ontario, Canada.
Organizational Affiliation: