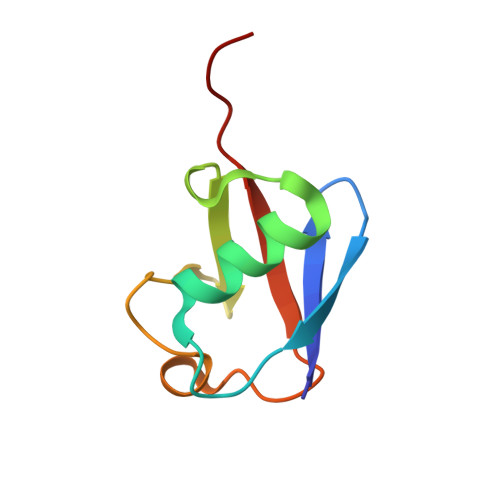
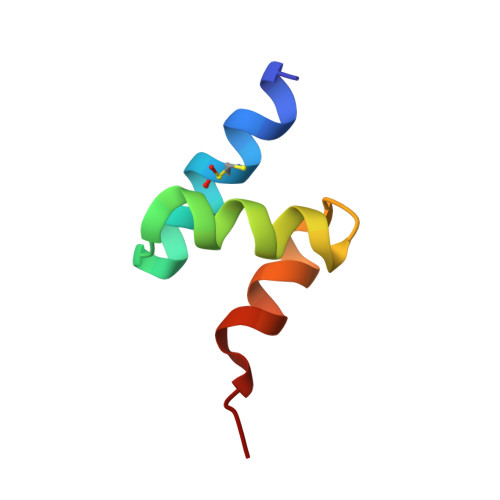
Ubiquitin stimulated reversal of topoisomerase 2 DNA-protein crosslinks by TDP2.
Schellenberg, M.J., Appel, C.D., Riccio, A.A., Butler, L.R., Krahn, J.M., Liebermann, J.A., Cortes-Ledesma, F., Williams, R.S.(2020) Nucleic Acids Res 48: 6310-6325
- PubMed: 32356875 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkaa318
- Primary Citation Related Structures:
6Q00, 6Q01 - PubMed Abstract:
Tyrosyl-DNA phosphodiesterase 2 (TDP2) reverses Topoisomerase 2 DNA-protein crosslinks (TOP2-DPCs) in a direct-reversal pathway licensed by ZATTZNF451 SUMO2 E3 ligase and SUMOylation of TOP2. TDP2 also binds ubiquitin (Ub), but how Ub regulates TDP2 functions is unknown. Here, we show that TDP2 co-purifies with K63 and K27 poly-Ubiquitinated cellular proteins independently of, and separately from SUMOylated TOP2 complexes. Poly-ubiquitin chains of ≥ Ub3 stimulate TDP2 catalytic activity in nuclear extracts and enhance TDP2 binding of DNA-protein crosslinks in vitro. X-ray crystal structures and small-angle X-ray scattering analysis of TDP2-Ub complexes reveal that the TDP2 UBA domain binds K63-Ub3 in a 1:1 stoichiometric complex that relieves a UBA-regulated autoinhibitory state of TDP2. Our data indicates that that poly-Ub regulates TDP2-catalyzed TOP2-DPC removal, and TDP2 single nucleotide polymorphisms can disrupt the TDP2-Ubiquitin interface.
- Structural Cell Biology Group, Genome Integrity and Structural Biology Laboratory, National Institute of Environmental Health Sciences, NIH, US Department of Health and Human Services, Research Triangle Park, NC 27709, USA.
Organizational Affiliation: