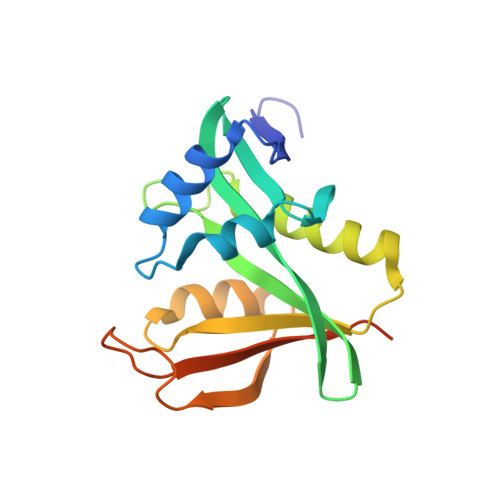
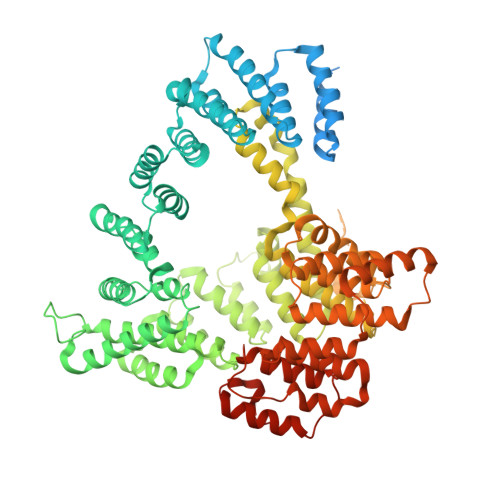
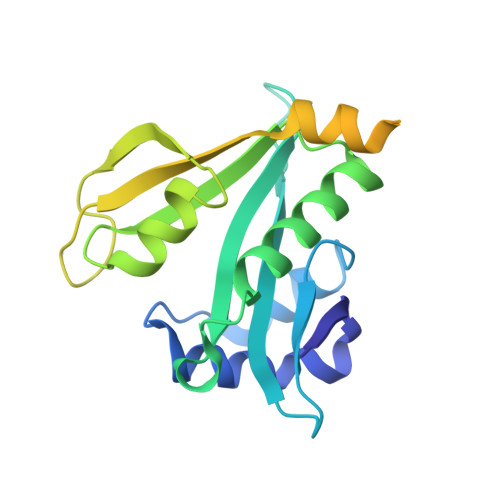
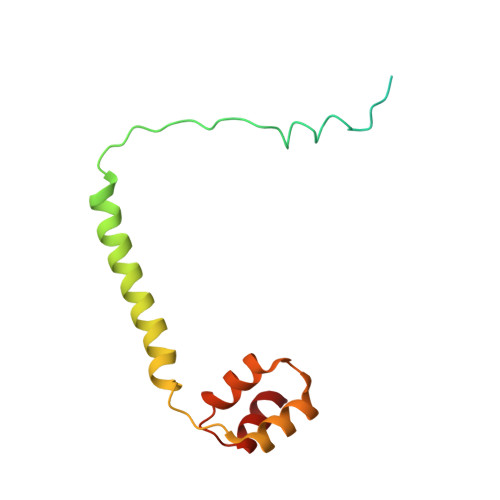
Molecular basis for N-terminal acetylation by human NatE and its modulation by HYPK.
Deng, S., McTiernan, N., Wei, X., Arnesen, T., Marmorstein, R.(2020) Nat Commun 11: 818-818
- PubMed: 32042062 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-14584-7
- Primary Citation Related Structures:
6PPL, 6PW9 - PubMed Abstract:
The human N-terminal acetyltransferase E (NatE) contains NAA10 and NAA50 catalytic, and NAA15 auxiliary subunits and associates with HYPK, a protein with intrinsic NAA10 inhibitory activity. NatE co-translationally acetylates the N-terminus of half the proteome to mediate diverse biological processes, including protein half-life, localization, and interaction. The molecular basis for how NatE and HYPK cooperate is unknown. Here, we report the cryo-EM structures of human NatE and NatE/HYPK complexes and associated biochemistry. We reveal that NAA50 and HYPK exhibit negative cooperative binding to NAA15 in vitro and in human cells by inducing NAA15 shifts in opposing directions. NAA50 and HYPK each contribute to NAA10 activity inhibition through structural alteration of the NAA10 substrate-binding site. NAA50 activity is increased through NAA15 tethering, but is inhibited by HYPK through structural alteration of the NatE substrate-binding site. These studies reveal the molecular basis for coordinated N-terminal acetylation by NatE and HYPK.
- Department of Chemistry, University of Pennsylvania, Philadelphia, PA, 19104, USA.
Organizational Affiliation: