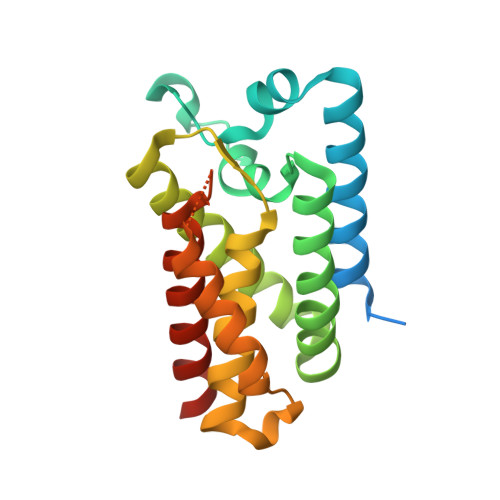
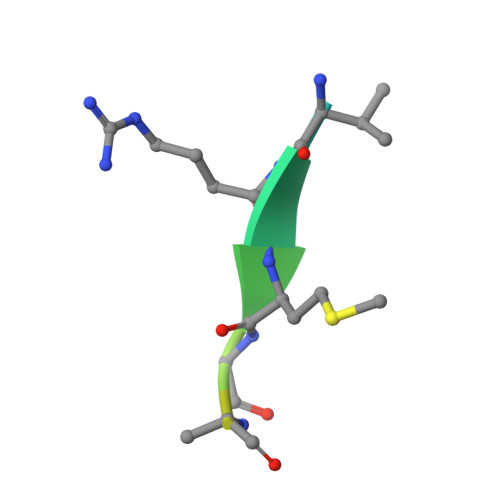
Ten catalytic snapshots of rhomboid intramembrane proteolysis from gate opening to peptide release.
Cho, S., Baker, R.P., Ji, M., Urban, S.(2019) Nat Struct Mol Biol 26: 910-918
- PubMed: 31570873 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-019-0296-9
- Primary Citation Related Structures:
6PJ4, 6PJ5, 6PJ7, 6PJ8, 6PJ9, 6PJA, 6PJP, 6PJQ, 6PJR, 6PJU - PubMed Abstract:
Protein cleavage inside the cell membrane triggers various pathophysiological signaling pathways, but the mechanism of catalysis is poorly understood. We solved ten structures of the Escherichia coli rhomboid protease in a bicelle membrane undergoing time-resolved steps that encompass the entire proteolytic reaction on a transmembrane substrate and an aldehyde inhibitor. Extensive gate opening accompanied substrate, but not inhibitor, binding, revealing that substrates and inhibitors take different paths to the active site. Catalysis unexpectedly commenced with, and was guided through subsequent catalytic steps by, motions of an extracellular loop, with local contributions from active site residues. We even captured the elusive tetrahedral intermediate that is uncleaved but covalently attached to the catalytic serine, about which the substrate was forced to bend dramatically. This unexpectedly stable intermediate indicates rhomboid catalysis uses an unprecedented reaction coordinate that may involve mechanically stressing the peptide bond, and could be selectively targeted by inhibitors.
- Department of Molecular Biology and Genetics, Johns Hopkins University School of Medicine, Baltimore, MD, USA.
Organizational Affiliation: