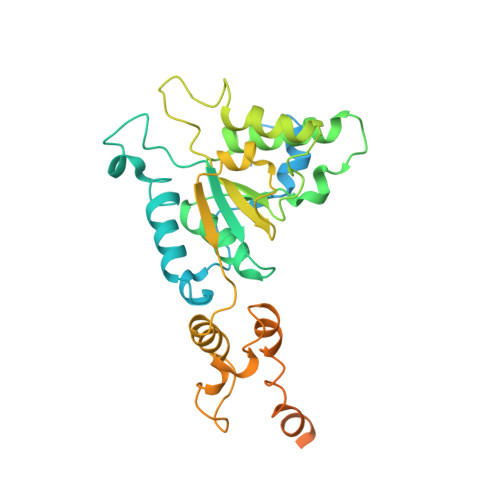
Structure of the AAA protein Msp1 reveals mechanism of mislocalized membrane protein extraction.
Wang, L., Myasnikov, A., Pan, X., Walter, P.(2020) Elife 9
- PubMed: 31999255 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.54031
- Primary Citation Related Structures:
6PDW, 6PDY, 6PE0 - PubMed Abstract:
The AAA protein Msp1 extracts mislocalized tail-anchored membrane proteins and targets them for degradation, thus maintaining proper cell organization. How Msp1 selects its substrates and firmly engages them during the energetically unfavorable extraction process remains a mystery. To address this question, we solved cryo-EM structures of Msp1-substrate complexes at near-atomic resolution. Akin to other AAA proteins, Msp1 forms hexameric spirals that translocate substrates through a central pore. A singular hydrophobic substrate recruitment site is exposed at the spiral's seam, which we propose positions the substrate for entry into the pore. There, a tight web of aromatic amino acids grips the substrate in a sequence-promiscuous, hydrophobic milieu. Elements at the intersubunit interfaces coordinate ATP hydrolysis with the subunits' positions in the spiral. We present a comprehensive model of Msp1's mechanism, which follows general architectural principles established for other AAA proteins yet specializes Msp1 for its unique role in membrane protein extraction.
- Howard Hughes Medical Institute, Chevy Chase, Maryland, United States.
Organizational Affiliation: