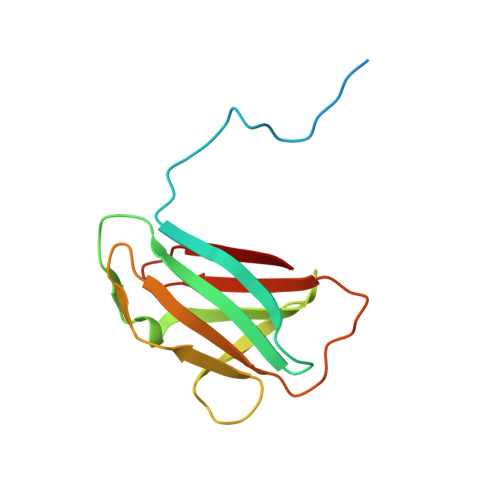
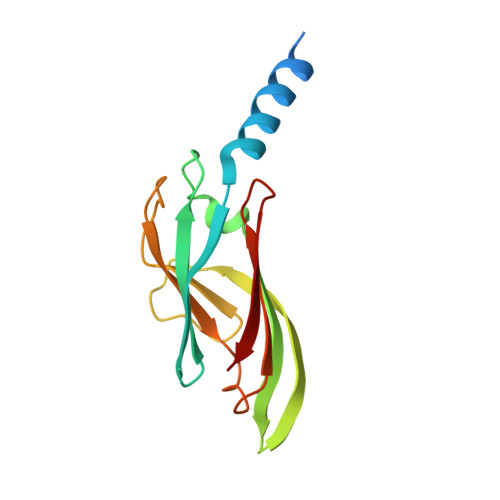
The structure of the periplasmic FlaG-FlaF complex and its essential role for archaellar swimming motility.
Tsai, C.L., Tripp, P., Sivabalasarma, S., Zhang, C., Rodriguez-Franco, M., Wipfler, R.L., Chaudhury, P., Banerjee, A., Beeby, M., Whitaker, R.J., Tainer, J.A., Albers, S.V.(2020) Nat Microbiol 5: 216-225
- PubMed: 31844299 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41564-019-0622-3
- Primary Citation Related Structures:
5TUG, 5TUH, 6PBK - PubMed Abstract:
Motility structures are vital in all three domains of life. In Archaea, motility is mediated by the archaellum, a rotating type IV pilus-like structure that is a unique nanomachine for swimming motility in nature. Whereas periplasmic FlaF binds the surface layer (S-layer), the structure, assembly and roles of other periplasmic components remain enigmatic, limiting our knowledge of the archaellum's functional interactions. Here, we find that the periplasmic protein FlaG and the association with its paralogue FlaF are essential for archaellation and motility. Therefore, we determine the crystal structure of Sulfolobus acidocaldarius soluble FlaG (sFlaG), which reveals a β-sandwich fold resembling the S-layer-interacting FlaF soluble domain (sFlaF). Furthermore, we solve the sFlaG 2 -sFlaF 2 co-crystal structure, define its heterotetrameric complex in solution by small-angle X-ray scattering and find that mutations that disrupt the complex abolish motility. Interestingly, the sFlaF and sFlaG of Pyrococcus furiosus form a globular complex, whereas sFlaG alone forms a filament, indicating that FlaF can regulate FlaG filament assembly. Strikingly, Sulfolobus cells that lack the S-layer component bound by FlaF assemble archaella but cannot swim. These collective results support a model where a FlaG filament capped by a FlaG-FlaF complex anchors the archaellum to the S-layer to allow motility.
- Department of Molecular and Cellular Oncology, The University of Texas MD Anderson Cancer Center, Houston, TX, USA.
Organizational Affiliation: