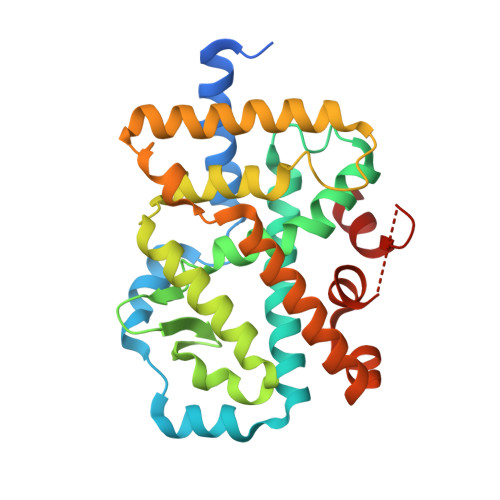
Identification of potent, selective and orally bioavailable phenyl ((R)-3-phenylpyrrolidin-3-yl)sulfone analogues as ROR gamma t inverse agonists.
Lu, Z., Duan, J.J., Xiao, H., Neels, J., Wu, D.R., Weigelt, C.A., Sack, J.S., Khan, J., Ruzanov, M., An, Y., Yarde, M., Karmakar, A., Vishwakrishnan, S., Baratam, V., Shankarappa, H., Vanteru, S., Babu, V., Basha, M., Kumar Gupta, A., Kumaravel, S., Mathur, A., Zhao, Q., Salter-Cid, L.M., Carter, P.H., Murali Dhar, T.G.(2019) Bioorg Med Chem Lett 29: 2265-2269
- PubMed: 31257087 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2019.06.036
- Primary Citation Related Structures:
6P9F - PubMed Abstract:
An X-ray crystal structure of one of our previously discovered RORγt inverse agonists bound to the RORγt ligand binding domain revealed that the cyclohexane carboxylic acid group of compound 2 plays a significant role in RORγt binding, forming four hydrogen bonding and ionic interactions with RORγt. SAR studies centered around the cyclohexane carboxylic acid group led to identification of several structurally diverse and more potent compounds, including new carboxylic acid analogues 7 and 20, and cyclic sulfone analogues 34 and 37. Notably, compounds 7 and 20 were found to maintain the desirable pharmacokinetic profile of 2.
- Research and Development, Bristol-Myers Squibb Company, Princeton, NJ 08543-4000, United States. Electronic address: zhonghui.lu@bms.com.
Organizational Affiliation: