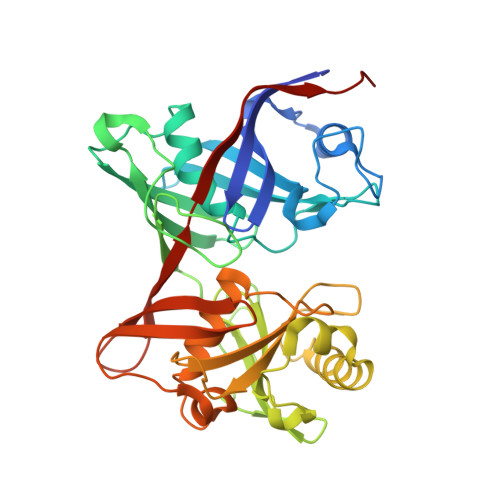
Structure and Chemical Reaction Mechanism of LigU, an Enzyme That Catalyzes an Allylic Isomerization in the Bacterial Degradation of Lignin.
Hogancamp, T.N., Cory, S.A., Barondeau, D.P., Raushel, F.M.(2019) Biochemistry 58: 3494-3503
- PubMed: 31339729 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.9b00549
- Primary Citation Related Structures:
6P3H, 6P3J, 6P3K - PubMed Abstract:
LigU from Novosphingobium sp. strain KA1 catalyzes the isomerization of (4 E )-oxalomesaconate (OMA) to (3 Z )-2-keto-4-carboxy-3-hexenedioate (KCH) as part of the protocatechuate (PCA) 4,5-cleavage pathway during the degradation of lignin. The three-dimensional structure of the apo form of the wild-type enzyme was determined by X-ray crystallography, and the structure of the K66M mutant enzyme was determined in the presence of the substrate OMA. LigU is a homodimer requiring no cofactors or metal ions with a diaminopimelate epimerase structural fold, consisting of two domains with similar topologies. Each domain has a central α-helix surrounded by a β-barrel composed of antiparallel β-strands. The active site is at the cleft of the two domains. 1 H nuclear magnetic resonance spectroscopy demonstrated that the enzyme catalyzes the exchange of the pro- S hydrogen at C5 of KCH with D 2 O during the isomerization reaction. Solvent-deuterium exchange experiments demonstrated that mutation of Lys-66 eliminated the isotope exchange at C5 and that mutation of C100 abolished exchange at C3. The positioning of these two residues in the active site of LigU is consistent with a reaction mechanism that is initiated by the abstraction of the pro- S hydrogen at C3 of OMA by the thiolate anion of Cys-100 and the donation of a proton at C5 of the proposed enolate anion intermediate by the side chain of Lys-66 to form the product KCH. The 1,3-proton transfer is suprafacial.
- Department of Chemistry , Texas A&M University , College Station , Texas 77843 , United States.
Organizational Affiliation: