A Human IgE Antibody Binding Site on Der p 2 for the Design of a Recombinant Allergen for Immunotherapy.
Glesner, J., Kapingidza, A.B., Godzwon, M., Offermann, L.R., Mueller, G.A., DeRose, E.F., Wright, P., Richardson, C.M., Woodfolk, J.A., Vailes, L.D., Wunschmann, S., London, R.E., Chapman, M.D., Ohlin, M., Chruszcz, M., Pomes, A.(2019) J Immunol 203: 2545-2556
- PubMed: 31554696 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.4049/jimmunol.1900580
- Primary Citation Related Structures:
6OY4 - PubMed Abstract:
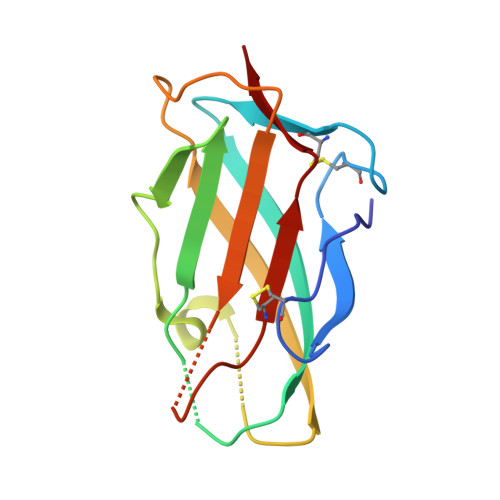
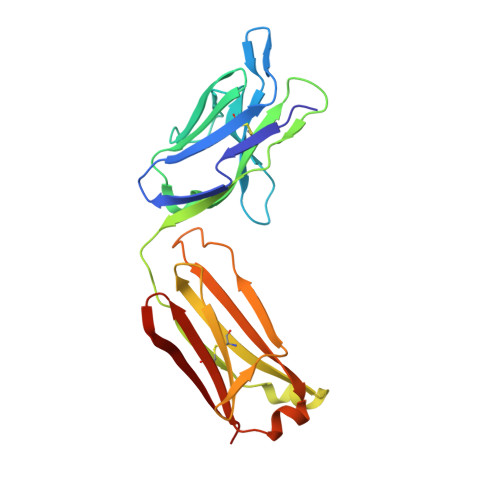
Der p 2 is one of the most important allergens from the house dust mite Dermatophagoides pteronyssinus Identification of human IgE Ab binding epitopes can be used for rational design of allergens with reduced IgE reactivity for therapy. Antigenic analysis of Der p 2 was performed by site-directed mutagenesis based on the x-ray crystal structure of the allergen in complex with a Fab from the murine IgG mAb 7A1 that binds an epitope overlapping with human IgE binding sites. Conformational changes upon Ab binding were confirmed by nuclear magnetic resonance using a 7A1-single-chain variable fragment. In addition, a human IgE Ab construct that interferes with mAb 7A1 binding was isolated from a combinatorial phage-display library constructed from a mite-allergic patient and expressed as two recombinant forms (single-chain Fab in Pichia pastoris and Fab in Escherichia coli ). These two IgE Ab constructs and the mAb 7A1 failed to recognize two Der p 2 epitope double mutants designed to abolish the allergen-Ab interaction while preserving the fold necessary to bind Abs at other sites of the allergen surface. A 10-100-fold reduction in binding of IgE from allergic subjects to the mutants additionally showed that the residues mutated were involved in IgE Ab binding. In summary, mutagenesis of a Der p 2 epitope defined by x-ray crystallography revealed an IgE Ab binding site that will be considered for the design of hypoallergens for immunotherapy.
- Indoor Biotechnologies, Inc., Charlottesville, VA 22903.
Organizational Affiliation: