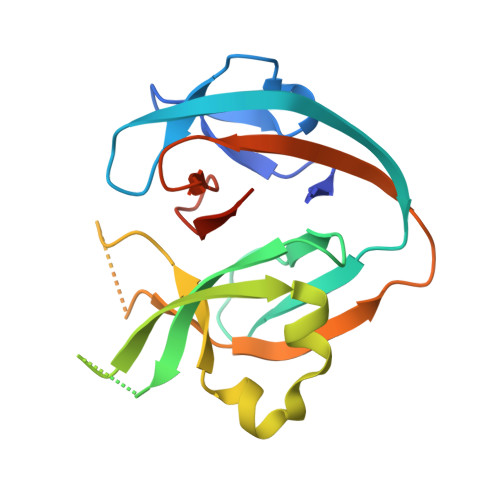
Spliceosomal Prp8 intein at the crossroads of protein and RNA splicing.
Green, C.M., Li, Z., Smith, A.D., Novikova, O., Bacot-Davis, V.R., Gao, F., Hu, S., Banavali, N.K., Thiele, D.J., Li, H., Belfort, M.(2019) PLoS Biol 17: e3000104-e3000104
- PubMed: 31600193 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pbio.3000104
- Primary Citation Related Structures:
6MX6, 6OWU - PubMed Abstract:
The spliceosome is a large ribonucleoprotein complex that removes introns from pre-mRNAs. At its functional core lies the essential pre-mRNA processing factor 8 (Prp8) protein. Across diverse eukaryotes, this protein cofactor of RNA catalysis harbors a self-splicing element called an intein. Inteins in Prp8 are extremely pervasive and are found at 7 different sites in various species. Here, we focus on the Prp8 intein from Cryptococcus neoformans (Cne), a human fungal pathogen. We solved the crystal structure of this intein, revealing structural homology among protein splicing sequences in eukaryotes, including the Hedgehog C terminus. Working with the Cne Prp8 intein in a reporter assay, we find that the biologically relevant divalent metals copper and zinc inhibit intein splicing, albeit by 2 different mechanisms. Copper likely stimulates reversible modifications on a catalytically important cysteine, whereas zinc binds at the terminal asparagine and the same critical cysteine. Importantly, we also show that copper treatment inhibits Prp8 protein splicing in Cne. Lastly, an intein-containing Prp8 precursor model is presented, suggesting that metal-induced protein splicing inhibition would disturb function of both Prp8 and the spliceosome. These results indicate that Prp8 protein splicing can be modulated, with potential functional implications for the spliceosome.
- Department of Biological Sciences and RNA Institute, University at Albany, Albany, New York, United States of America.
Organizational Affiliation: