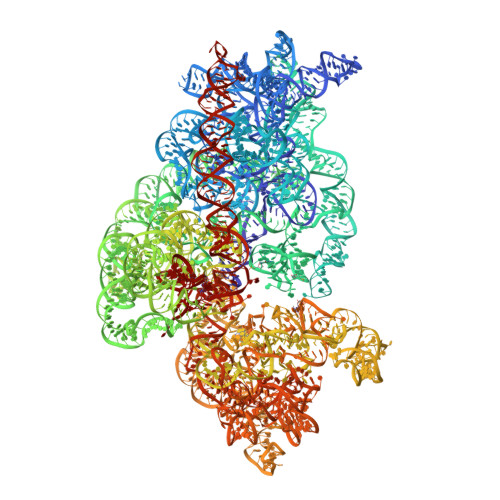
The structural basis for release-factor activation during translation termination revealed by time-resolved cryogenic electron microscopy.
Fu, Z., Indrisiunaite, G., Kaledhonkar, S., Shah, B., Sun, M., Chen, B., Grassucci, R.A., Ehrenberg, M., Frank, J.(2019) Nat Commun 10: 2579-2579
- PubMed: 31189921 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-10608-z
- Primary Citation Related Structures:
6ORE, 6ORL, 6OSK, 6OSQ, 6OST, 6OT3, 6OUO - PubMed Abstract:
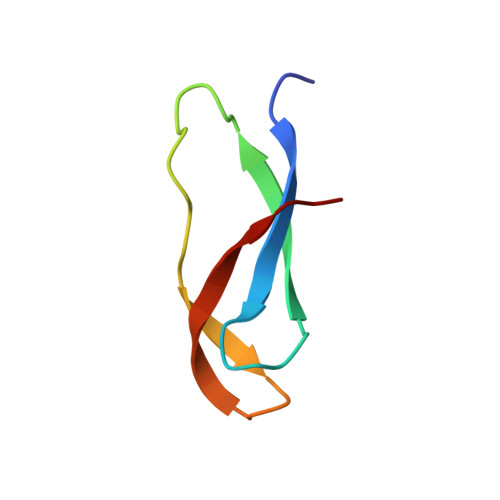
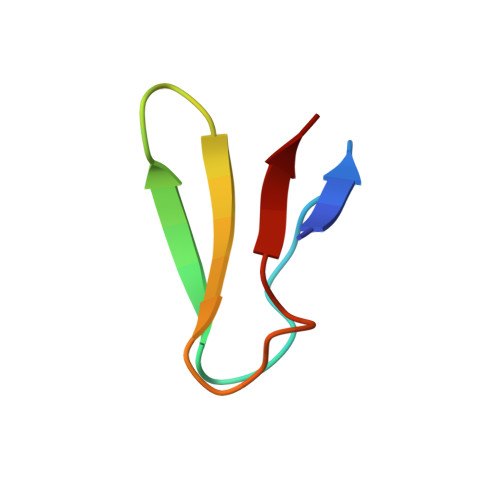
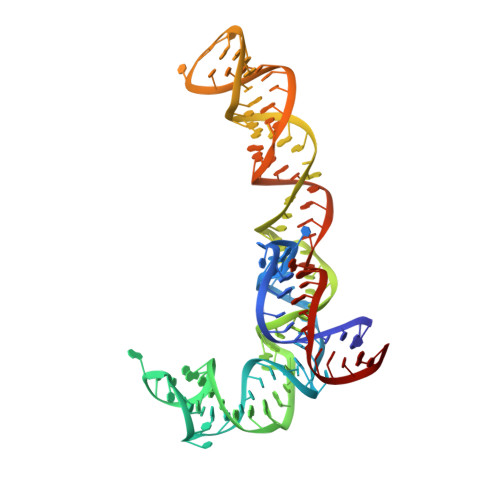
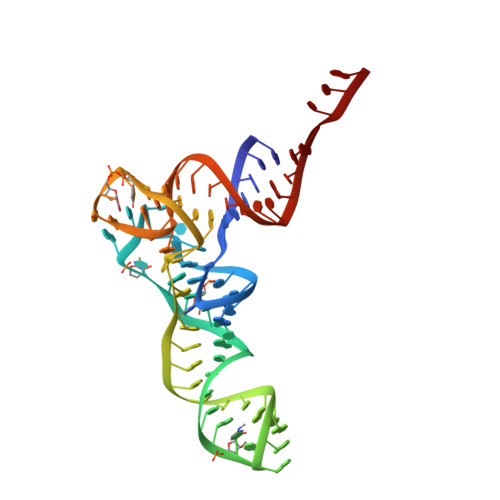
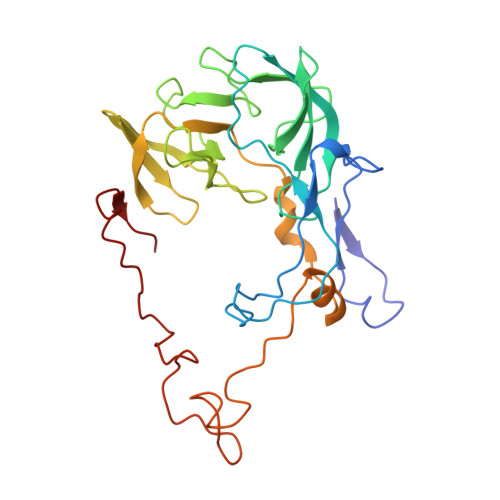
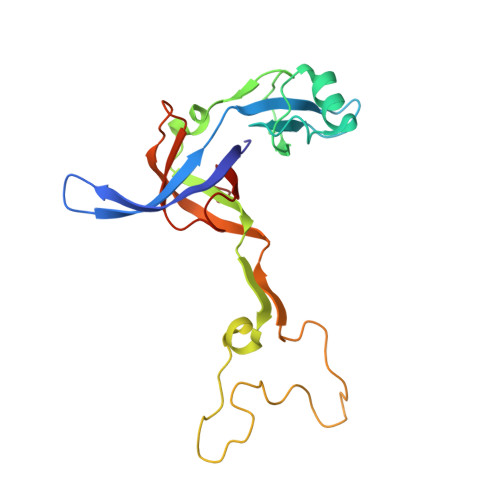
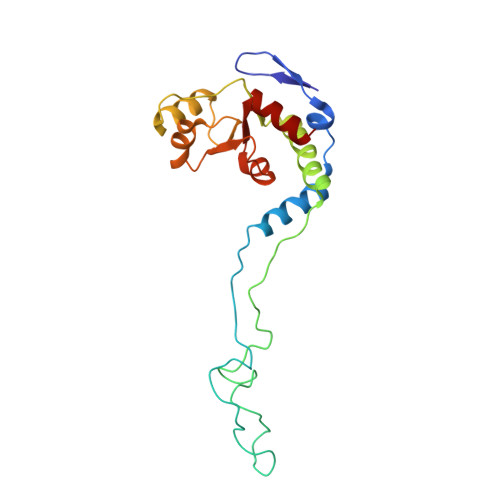
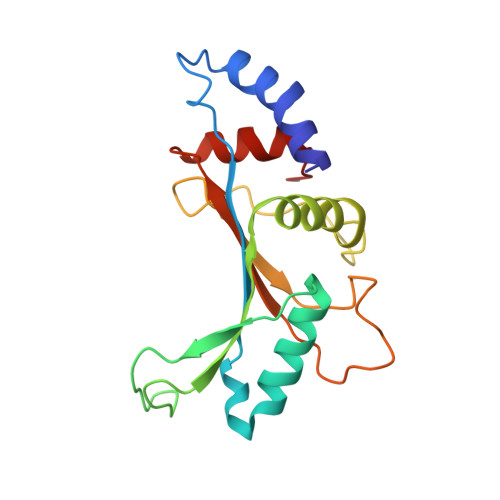
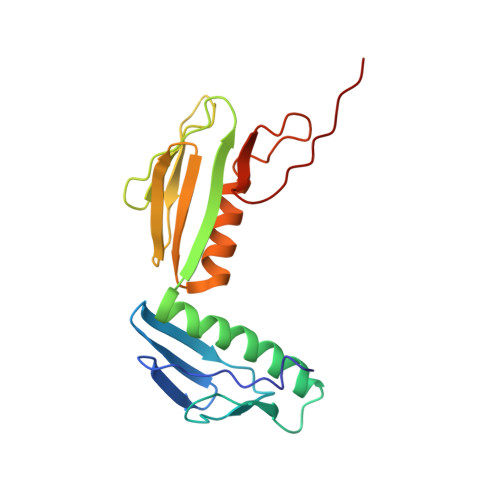
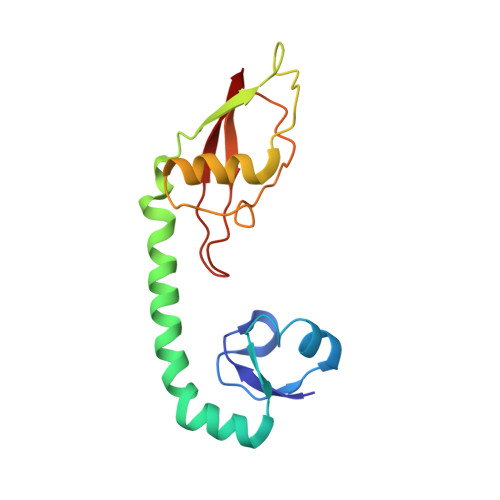
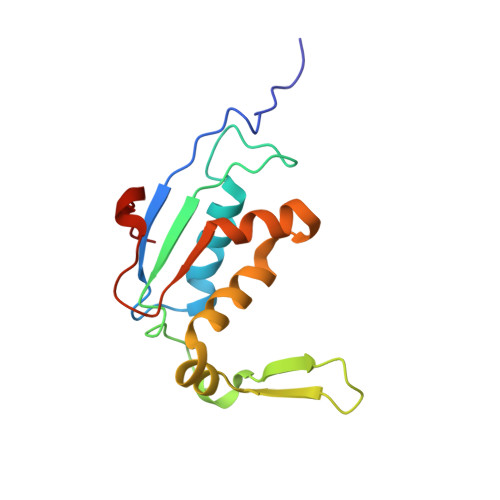
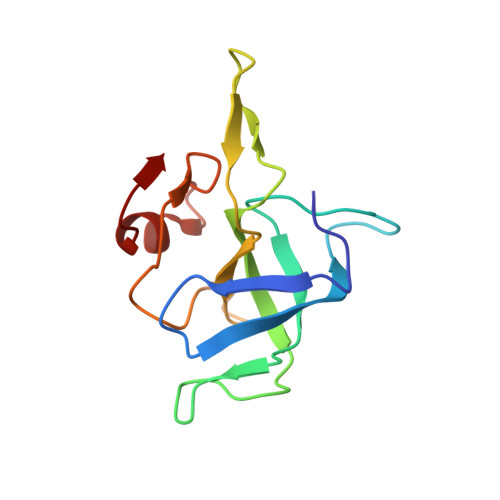
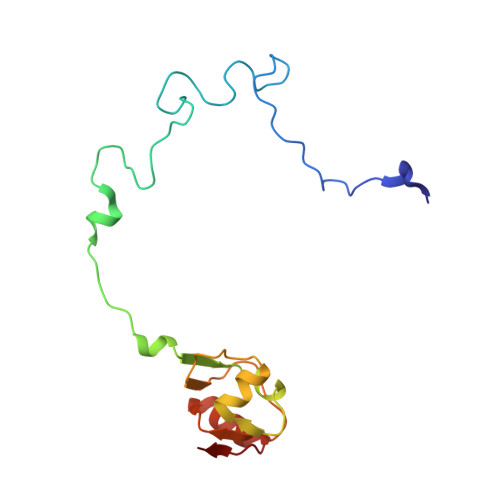
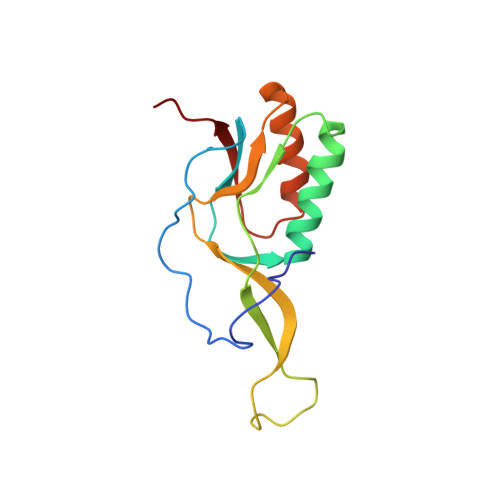
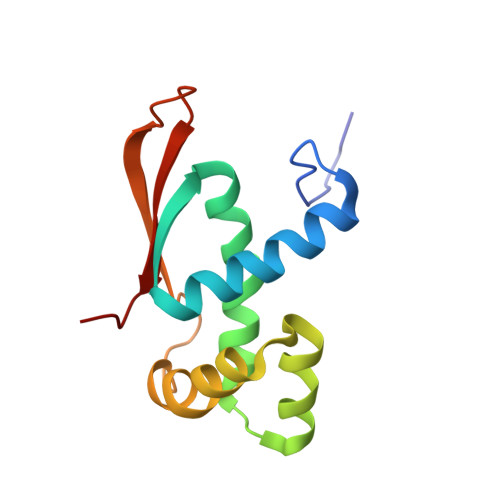
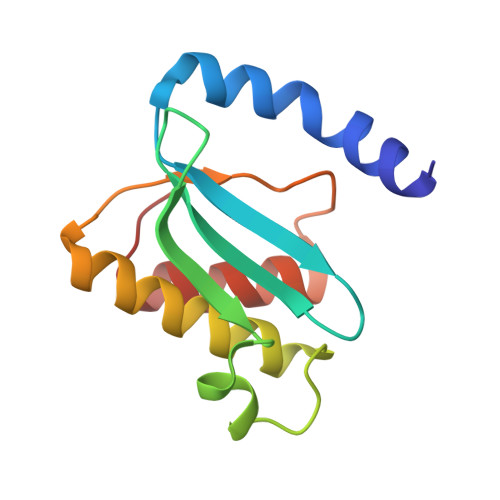
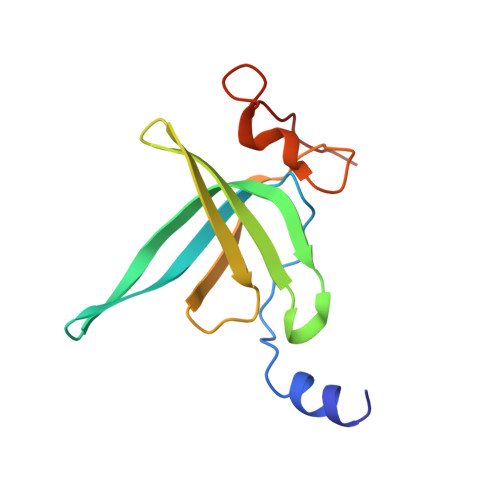
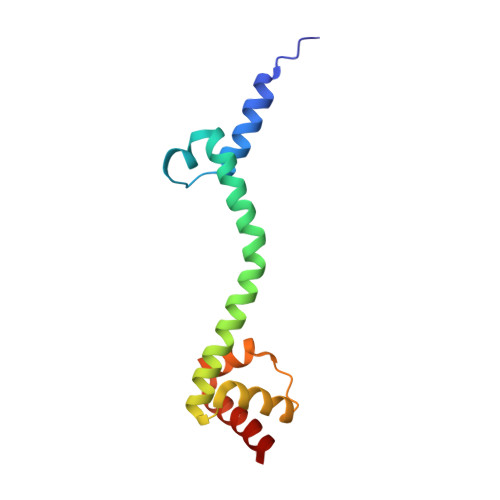
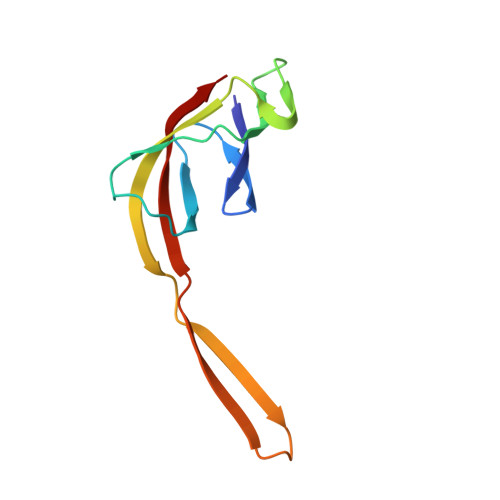
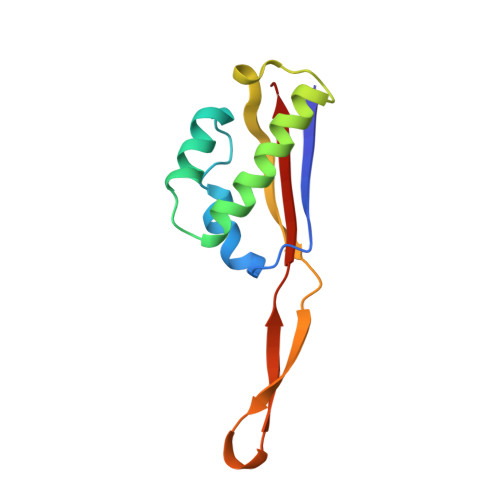
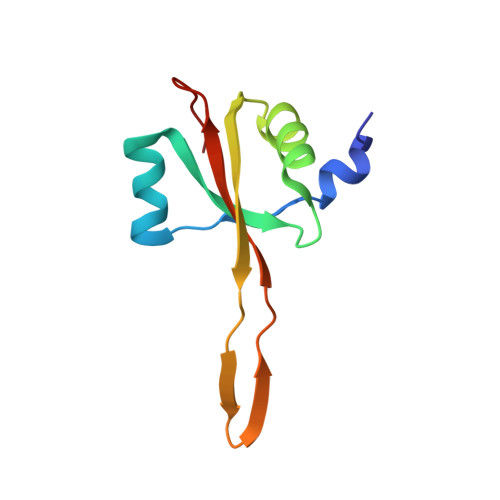
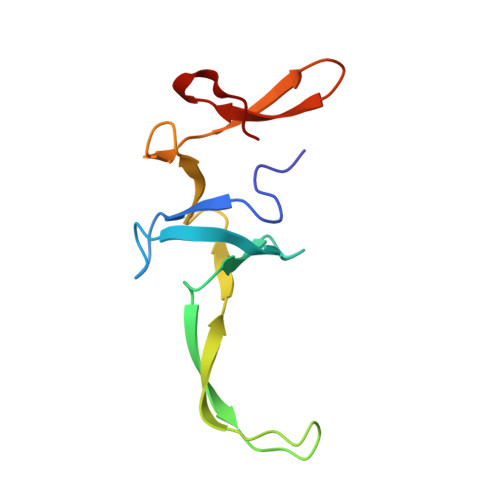
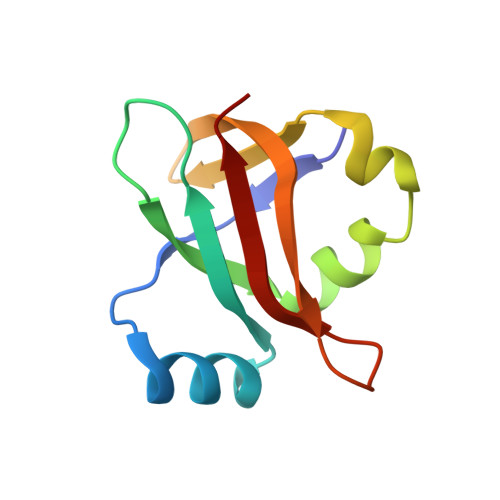
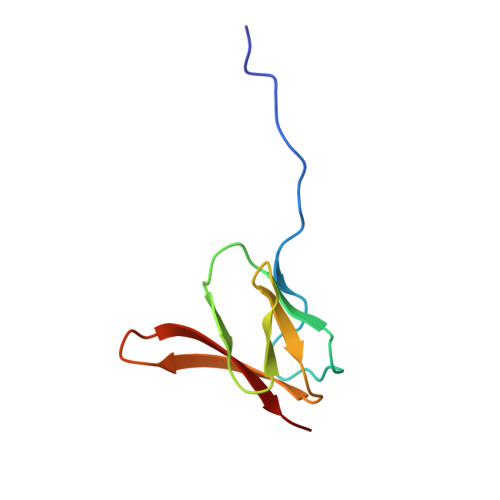
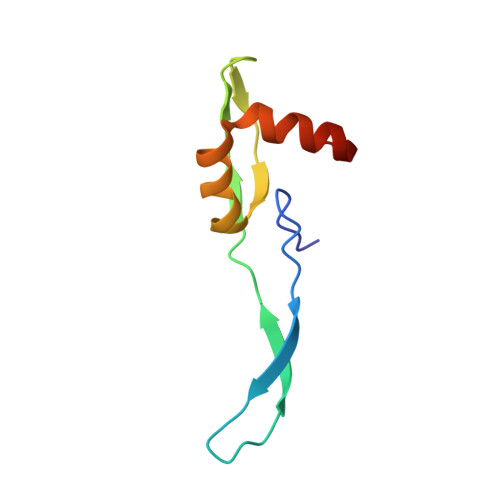
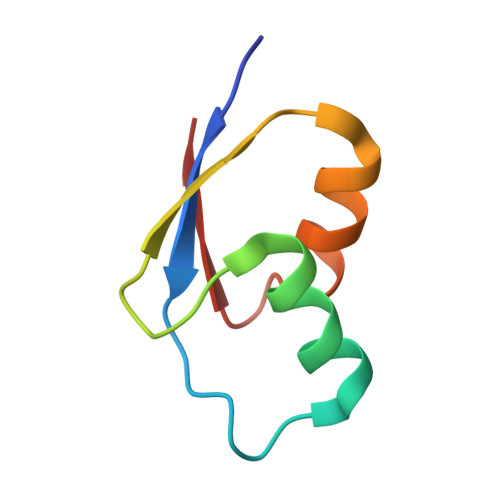
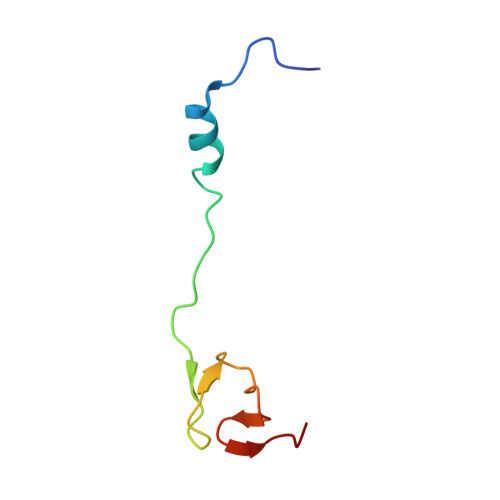
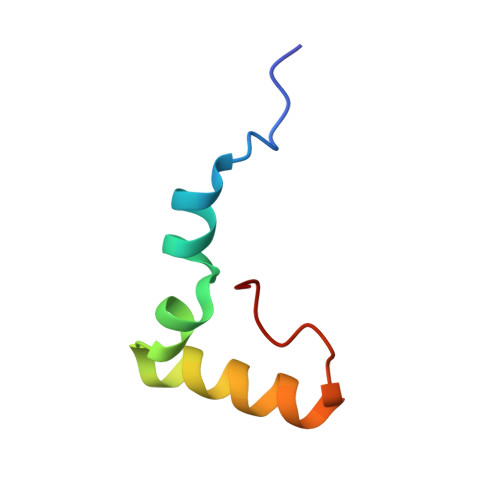
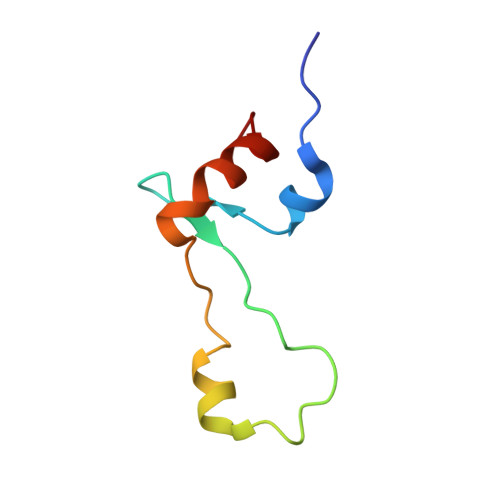
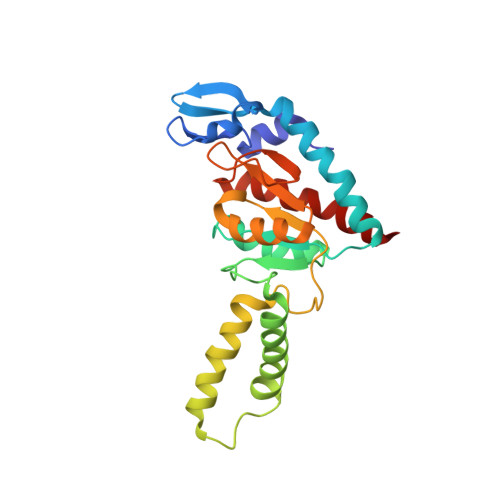
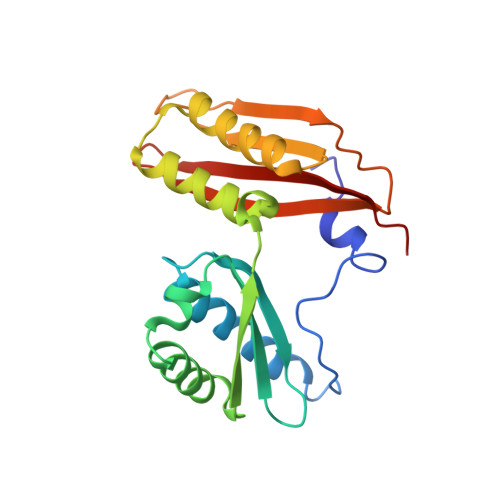
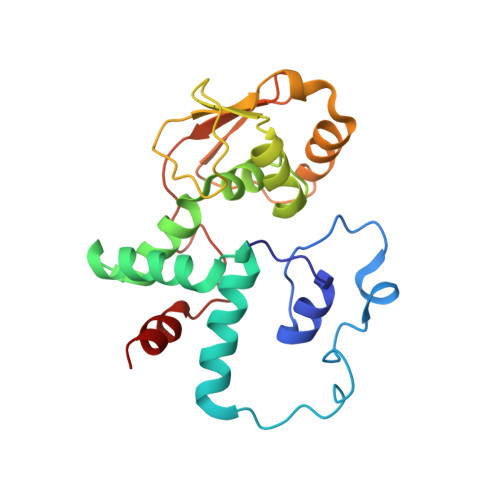
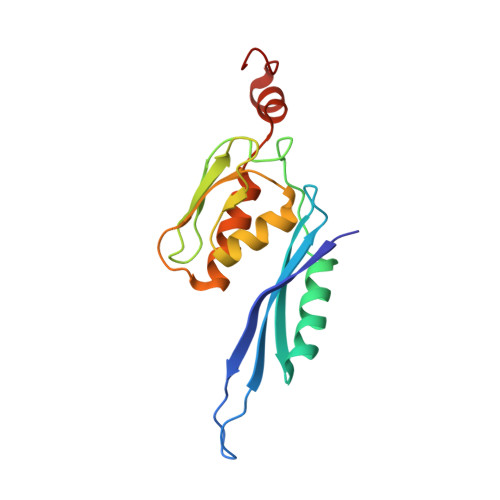
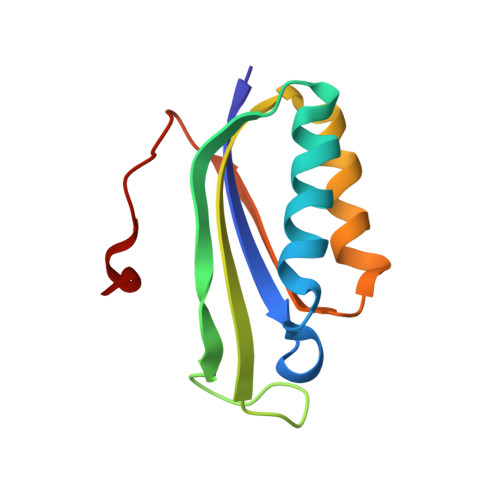
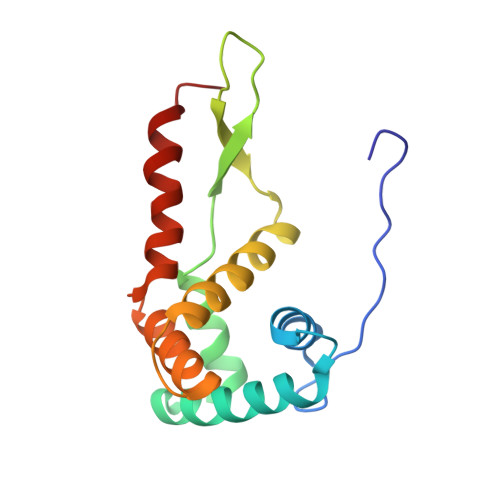
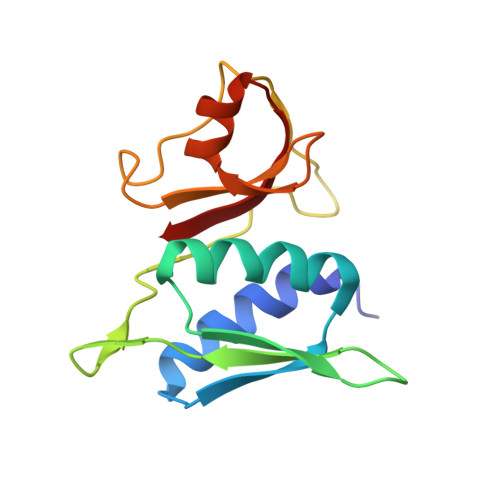
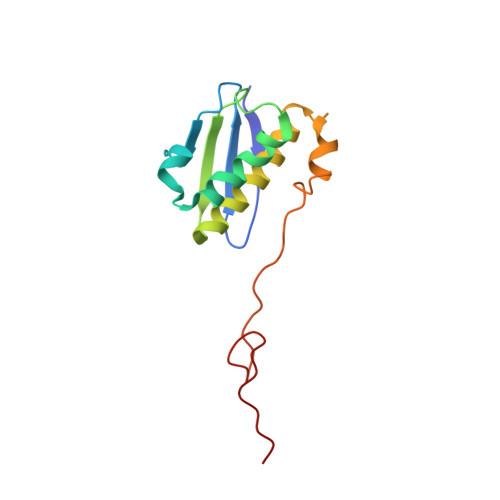
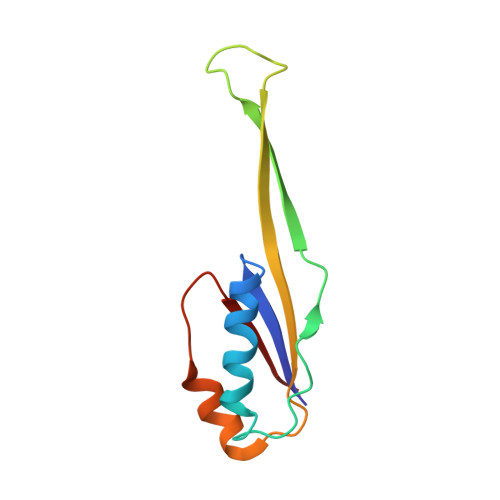
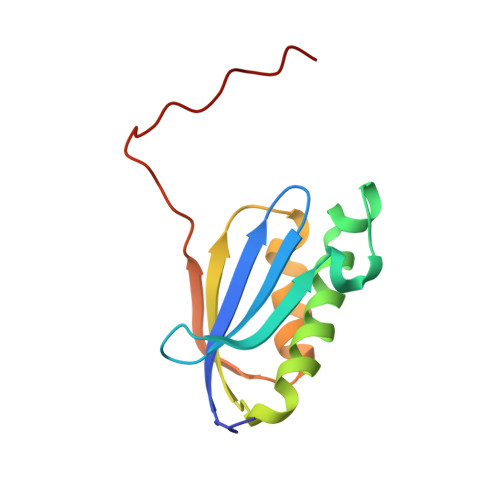
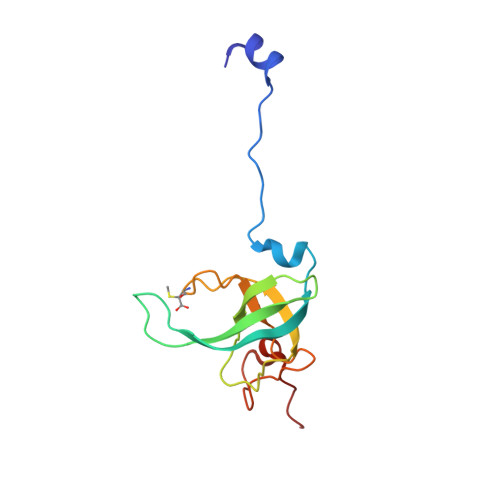
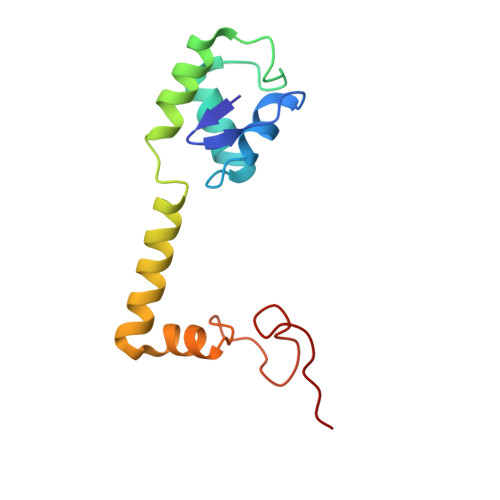
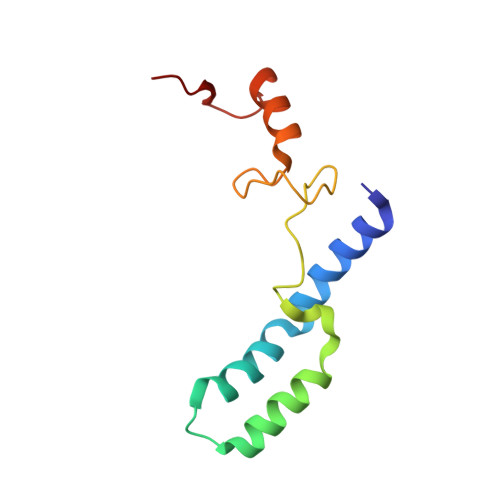
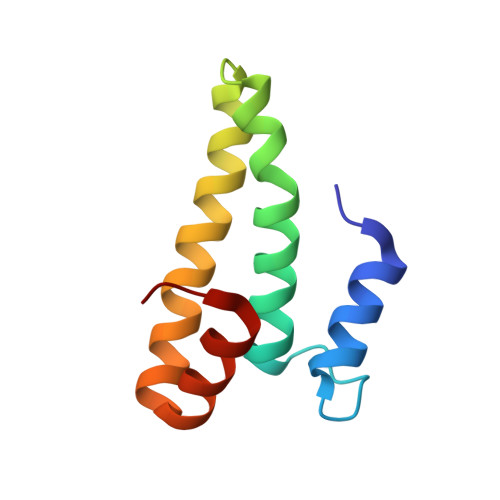
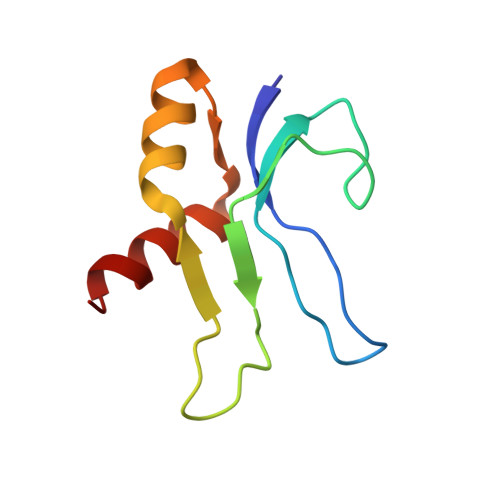
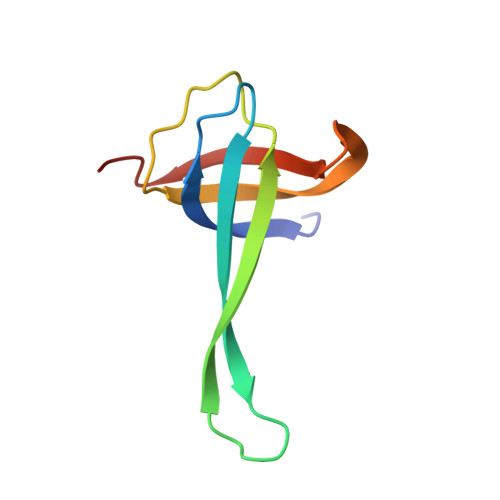
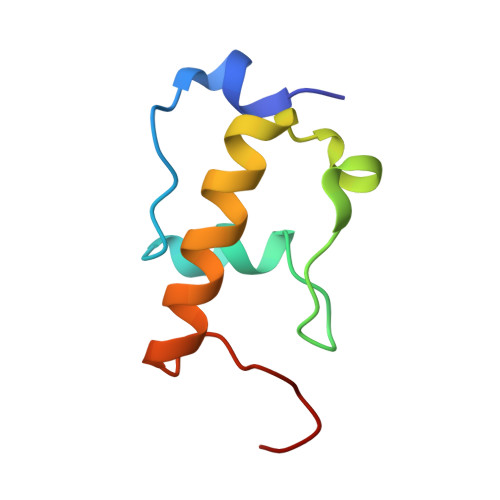
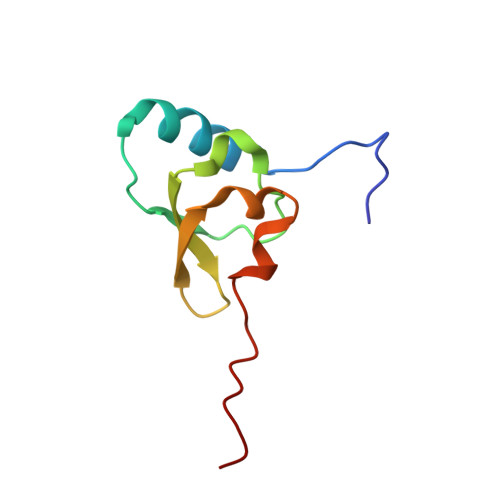
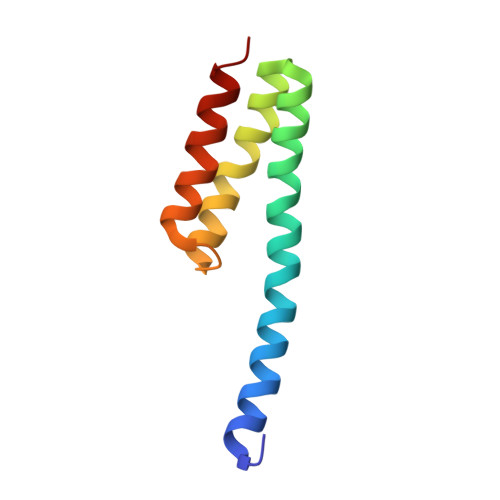
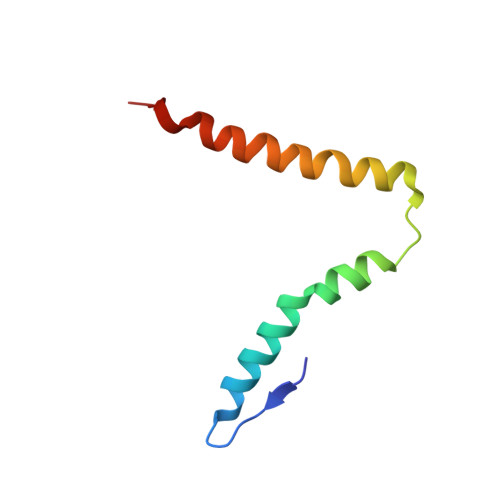
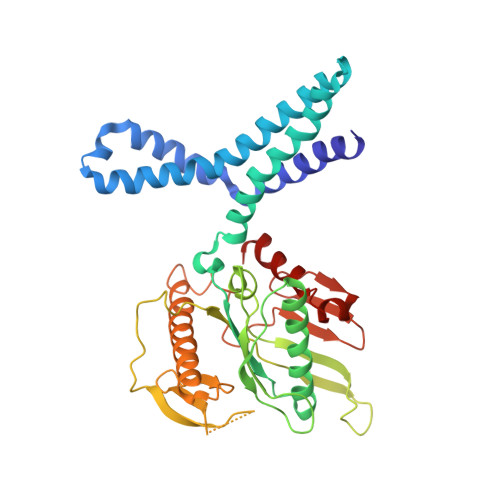
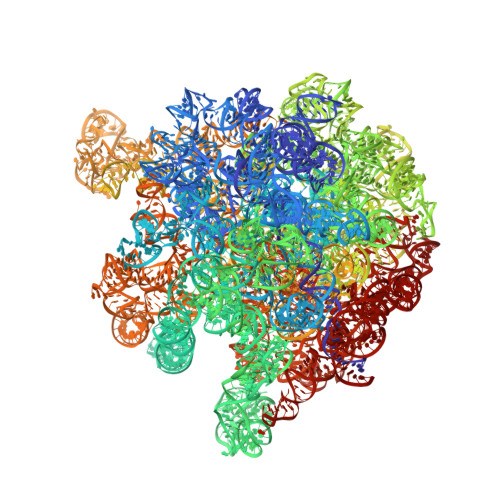
When the ribosome encounters a stop codon, it recruits a release factor (RF) to hydrolyze the ester bond between the peptide chain and tRNA. RFs have structural motifs that recognize stop codons in the decoding center and a GGQ motif for induction of hydrolysis in the peptidyl transfer center 70 Å away. Surprisingly, free RF2 is compact, with only 20 Å between its codon-reading and GGQ motifs. Cryo-EM showed that ribosome-bound RFs have extended structures, suggesting that RFs are compact when entering the ribosome and then extend their structures upon stop codon recognition. Here we use time-resolved cryo-EM to visualize transient compact forms of RF1 and RF2 at 3.5 and 4 Å resolution, respectively, in the codon-recognizing ribosome complex on the native pathway. About 25% of complexes have RFs in the compact state at 24 ms reaction time, and within 60 ms virtually all ribosome-bound RFs are transformed to their extended forms.
- Department of Biochemistry and Molecular Biophysics, Columbia University, New York, NY, 10032, USA.
Organizational Affiliation: