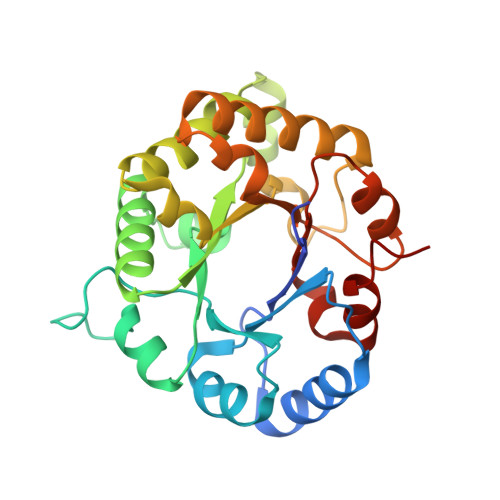
Crystal structures of Triosephosphate Isomerases from Taenia solium and Schistosoma mansoni provide insights for vaccine rationale and drug design against helminth parasites.
Jimenez-Sandoval, P., Castro-Torres, E., Gonzalez-Gonzalez, R., Diaz-Quezada, C., Gurrola, M., Camacho-Manriquez, L.D., Leyva-Navarro, L., Brieba, L.G.(2020) PLoS Negl Trop Dis 14: e0007815-e0007815
- PubMed: 31923219 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pntd.0007815
- Primary Citation Related Structures:
6OOG, 6OOI - PubMed Abstract:
Triosephosphate isomerases (TPIs) from Taenia solium (TsTPI) and Schistosoma mansoni (SmTPI) are potential vaccine and drug targets against cysticercosis and schistosomiasis, respectively. This is due to the dependence of parasitic helminths on glycolysis and because those proteins elicit an immune response, presumably due to their surface localization. Here we report the crystal structures of TsTPI and SmTPI in complex with 2-phosphoglyceric acid (2-PGA). Both TPIs fold into a dimeric (β-α)8 barrel in which the dimer interface consists of α-helices 2, 3, and 4, and swapping of loop 3. TPIs from parasitic helminths harbor a region of three amino acids knows as the SXD/E insert (S155 to E157 and S157 to D159 in TsTPI and SmTPI, respectively). This insert is located between α5 and β6 and is proposed to be the main TPI epitope. This region is part of a solvent-exposed 310-helix that folds into a hook-like structure. The crystal structures of TsTPI and SmTPI predicted conformational epitopes that could be used for vaccine design. Surprisingly, the epitopes corresponding to the SXD/E inserts are not the ones with the greatest immunological potential. SmTPI, but not TsTPI, habors a sole solvent exposed cysteine (SmTPI-S230) and alterations in this residue decrease catalysis. The latter suggests that thiol-conjugating agents could be used to target SmTPI. In sum, the crystal structures of SmTPI and TsTPI are a blueprint for targeted schistosomiasis and cysticercosis drug and vaccine development.
- Laboratorio Nacional de Genómica para la Biodiversidad, Centro de Investigación y de Estudios Avanzados del IPN, Irapuato, Guanajuato, México.
Organizational Affiliation: