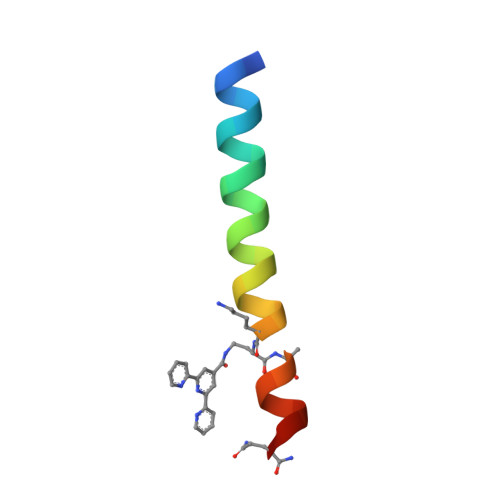
Understanding and controlling the metal-directed assembly of terpyridine-functionalized coiled-coil peptides.
Scheib, K.A., Tavenor, N.A., Lawless, M.J., Saxena, S., Horne, W.S.(2019) Chem Commun (Camb) 55: 7752-7755
- PubMed: 31204733 Search on PubMed
- DOI: https://doi.org/10.1039/c9cc03496j
- Primary Citation Related Structures:
6OLN, 6OLO - PubMed Abstract:
Metal-binding peptides are versatile building blocks in supramolecular chemistry. We recently reported a class of crystalline materials formed through a combination of coiled-coil peptide self-association and metal coordination. Here, we probe the serendipitously discovered metal binding motif that drives the assembly and apply these insights to exert rational control over structure and morphology in the materials.
- Department of Chemistry, University of Pittsburgh, Pittsburgh, PA 15260, USA. horne@pitt.edu.
Organizational Affiliation: