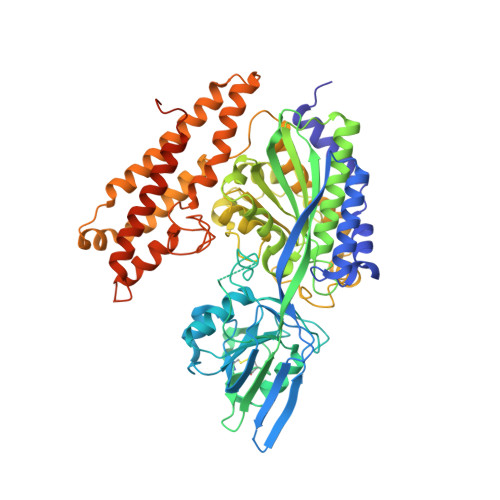
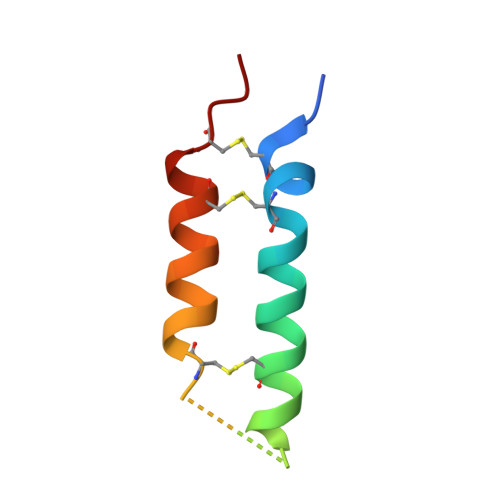
A TfR-Binding Cystine-Dense Peptide Promotes Blood-Brain Barrier Penetration of Bioactive Molecules.
Crook, Z.R., Girard, E., Sevilla, G.P., Merrill, M., Friend, D., Rupert, P.B., Pakiam, F., Nguyen, E., Yin, C., Ruff, R.O., Hopping, G., Strand, A.D., Finton, K.A.K., Coxon, M., Mhyre, A.J., Strong, R.K., Olson, J.M.(2020) J Mol Biology 432: 3989-4009
- PubMed: 32304700 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2020.04.002
- Primary Citation Related Structures:
6OKD, 6OKE - PubMed Abstract:
The impenetrability of the blood-brain barrier (BBB) to most conventional drugs impedes the treatment of central nervous system (CNS) disorders. Interventions for diseases like brain cancer, neurodegeneration, or age-associated inflammatory processes require varied approaches to CNS drug delivery. Cystine-dense peptides (CDPs) have drawn recent interest as drugs or drug-delivery vehicles. Found throughout the phylogenetic tree, often in drug-like roles, their size, stability, and protein interaction capabilities make CDPs an attractive mid-size biologic scaffold to complement conventional antibody-based drugs. Here, we describe the identification, maturation, characterization, and utilization of a CDP that binds to the transferrin receptor (TfR), a native receptor and BBB transporter for the iron chaperone transferrin. We developed variants with varying binding affinities (K D as low as 216 pM), co-crystallized it with the receptor, and confirmed murine cross-reactivity. It accumulates in the mouse CNS at ~25% of blood levels (CNS blood content is only ~1%-6%) and delivers neurotensin, an otherwise non-BBB-penetrant neuropeptide, at levels capable of modulating CREB signaling in the mouse brain. Our work highlights the utility of CDPs as a diverse, easy-to-screen scaffold family worthy of inclusion in modern drug discovery strategies, demonstrated by the discovery of a candidate CNS drug delivery vehicle ready for further optimization and preclinical development.
- Clinical Research Division, Fred Hutchinson Cancer Research Center, 1100 Fairview Ave N, Seattle, WA 98109, USA.
Organizational Affiliation: