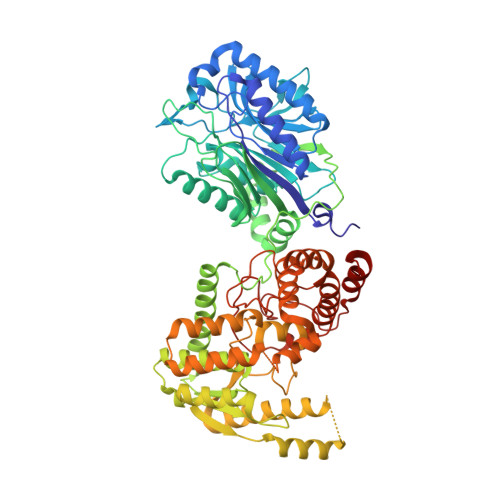
Different ways to transport ammonia in human and Mycobacterium tuberculosis NAD+synthetases.
Chuenchor, W., Doukov, T.I., Chang, K.T., Resto, M., Yun, C.S., Gerratana, B.(2020) Nat Commun 11: 16-16
- PubMed: 31911602 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-13845-4
- Primary Citation Related Structures:
6OFB, 6OFC - PubMed Abstract:
NAD + synthetase is an essential enzyme of de novo and recycling pathways of NAD + biosynthesis in Mycobacterium tuberculosis but not in humans. This bifunctional enzyme couples the NAD + synthetase and glutaminase activities through an ammonia tunnel but free ammonia is also a substrate. Here we show that the Homo sapiens NAD + synthetase (hsNadE) lacks substrate specificity for glutamine over ammonia and displays a modest activation of the glutaminase domain compared to tbNadE. We report the crystal structures of hsNadE and NAD + synthetase from M. tuberculosis (tbNadE) with synthetase intermediate analogues. Based on the observed exclusive arrangements of the domains and of the intra- or inter-subunit tunnels we propose a model for the inter-domain communication mechanism for the regulation of glutamine-dependent activity and NH 3 transport. The structural and mechanistic comparison herein reported between hsNadE and tbNadE provides also a starting point for future efforts in the development of anti-TB drugs.
- Departments of Chemistry and Biochemistry, University of Maryland, College Park, MD, 20742, USA.
Organizational Affiliation: