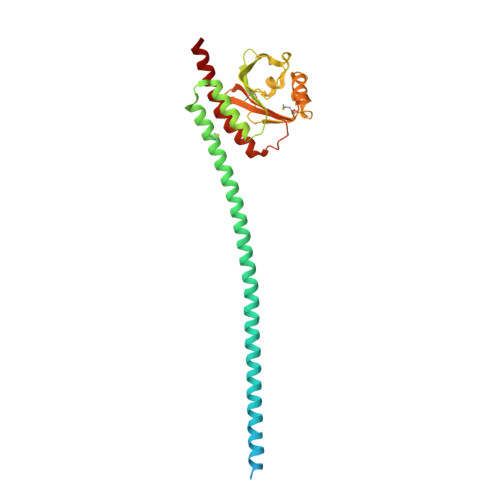
Structure and function of Spc42 coiled-coils in yeast centrosome assembly and duplication.
Drennan, A.C., Krishna, S., Seeger, M.A., Andreas, M.P., Gardner, J.M., Sether, E.K.R., Jaspersen, S.L., Rayment, I.(2019) Mol Biol Cell 30: 1505-1522
- PubMed: 30969903 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1091/mbc.E19-03-0167
- Primary Citation Related Structures:
6OD2, 6OEC, 6OEI - PubMed Abstract:
Centrosomes and spindle pole bodies (SPBs) are membraneless organelles whose duplication and assembly is necessary for bipolar mitotic spindle formation. The structural organization and functional roles of major proteins in these organelles can provide critical insights into cell division control. Spc42, a phosphoregulated protein with an N-terminal dimeric coiled-coil (DCC), assembles into a hexameric array at the budding yeast SPB core, where it functions as a scaffold for SPB assembly. Here, we present in vitro and in vivo data to elucidate the structural arrangement and biological roles of Spc42 elements. Crystal structures reveal details of two additional coiled-coils in Spc42: a central trimeric coiled-coil and a C-terminal antiparallel DCC. Contributions of the three Spc42 coiled-coils and adjacent undetermined regions to the formation of an ∼145 Å hexameric lattice in an in vitro lipid monolayer assay and to SPB duplication and assembly in vivo reveal structural and functional redundancy in Spc42 assembly. We propose an updated model that incorporates the inherent symmetry of these Spc42 elements into a lattice, and thereby establishes the observed sixfold symmetry. The implications of this model for the organization of the central SPB core layer are discussed.
- Department of Biochemistry, University of Wisconsin-Madison, WI 53706.
Organizational Affiliation: