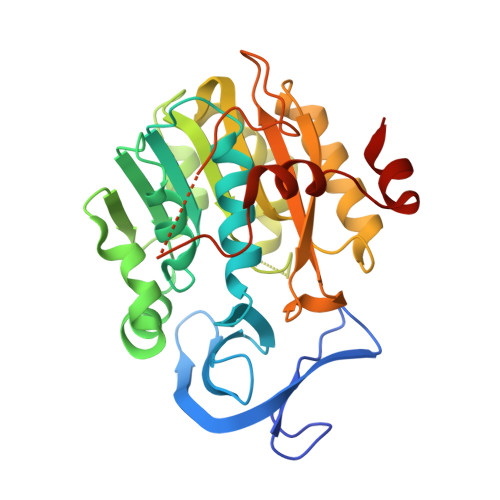
Spermidine Synthase (SPDS) Undergoes Concerted Structural Rearrangements Upon Ligand Binding - A Case Study of the Two SPDS Isoforms FromArabidopsis thaliana.
Sekula, B., Dauter, Z.(2019) Front Plant Sci 10: 555-555
- PubMed: 31134111 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/fpls.2019.00555
- Primary Citation Related Structures:
6O63, 6O64, 6O65 - PubMed Abstract:
Spermidine synthases (SPDSs) catalyze the production of the linear triamine, spermidine, from putrescine. They utilize decarboxylated S-adenosylmethionine (dc-SAM), a universal cofactor of aminopropyltransferases, as a donor of the aminopropyl moiety. In this work, we describe crystal structures of two SPDS isoforms from Arabidopsis thaliana ( At SPDS1 and At SPDS2). At SPDS1 and At SPDS2 are dimeric enzymes that share the fold of the polyamine biosynthesis proteins. Subunits of both isoforms present the characteristic two-domain structure. Smaller, N-terminal domain is built of the two β-sheets, while the C-terminal domain has a Rossmann fold-like topology. The catalytic cleft composed of two main compartments, the dc-SAM binding site and the polyamine groove, is created independently in each At SPDS subunits at the domain interface. We also provide the structural details about the dc-SAM binding mode and the inhibition of SPDS by a potent competitive inhibitor, cyclohexylamine (CHA). CHA occupies the polyamine binding site of At SPDS where it is bound at the bottom of the active site with the amine group placed analogously to the substrate. The crystallographic snapshots show in detail the structural rearrangements of At SPDS1 and At SPDS2 that are required to stabilize ligands within the active site. The concerted movements are observed in both compartments of the catalytic cleft, where three major parts significantly change their conformation. These are (i) the neighborhood of the glycine-rich region where aminopropyl moiety of dc-SAM is bound, (ii) the very flexible gate region with helix η6, which interacts with both, the adenine moiety of dc-SAM and the bound polyamine or inhibitor, and (iii) the N-terminal β-hairpin, that limits the putrescine binding grove at the bottom of the catalytic site.
- Synchrotron Radiation Research Section, Macromolecular Crystallography Laboratory, National Cancer Institute, Argonne, IL, United States.
Organizational Affiliation: