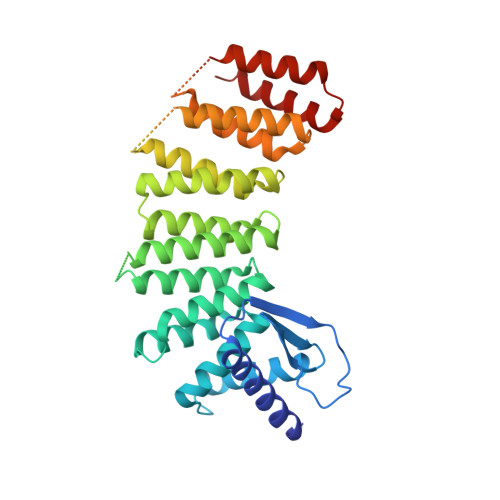
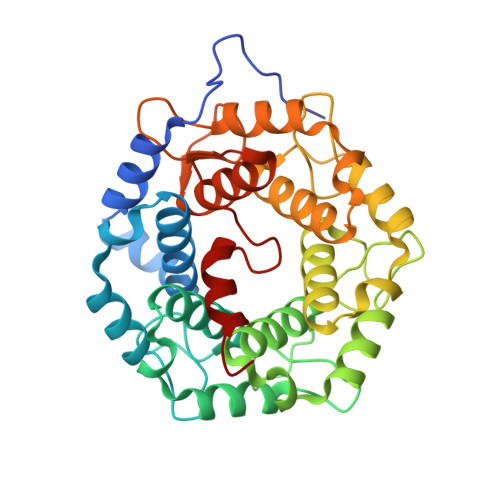
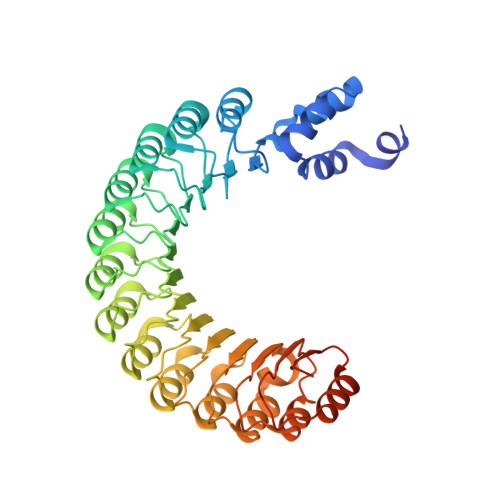
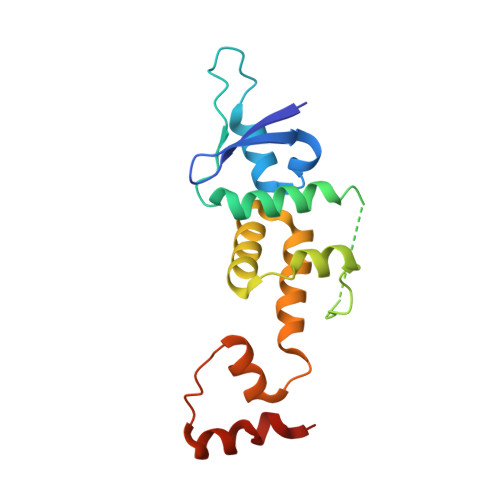
GGTase3 is a newly identified geranylgeranyltransferase targeting a ubiquitin ligase.
Kuchay, S., Wang, H., Marzio, A., Jain, K., Homer, H., Fehrenbacher, N., Philips, M.R., Zheng, N., Pagano, M.(2019) Nat Struct Mol Biol 26: 628-636
- PubMed: 31209342 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-019-0249-3
- Primary Citation Related Structures:
6O60 - PubMed Abstract:
Protein prenylation is believed to be catalyzed by three heterodimeric enzymes: FTase, GGTase1 and GGTase2. Here we report the identification of a previously unknown human prenyltransferase complex consisting of an orphan prenyltransferase α-subunit, PTAR1, and the catalytic β-subunit of GGTase2, RabGGTB. This enzyme, which we named GGTase3, geranylgeranylates FBXL2 to allow its localization at cell membranes, where this ubiquitin ligase mediates the polyubiquitylation of membrane-anchored proteins. In cells, FBXL2 is specifically recognized by GGTase3 despite having a typical carboxy-terminal CaaX prenylation motif that is predicted to be recognized by GGTase1. Our crystal structure analysis of the full-length GGTase3-FBXL2-SKP1 complex reveals an extensive multivalent interface specifically formed between the leucine-rich repeat domain of FBXL2 and PTAR1, which unmasks the structural basis of the substrate-enzyme specificity. By uncovering a missing prenyltransferase and its unique mode of substrate recognition, our findings call for a revision of the 'prenylation code'.
- Department of Biochemistry and Molecular Pharmacology, New York University School of Medicine, New York, NY, USA.
Organizational Affiliation: