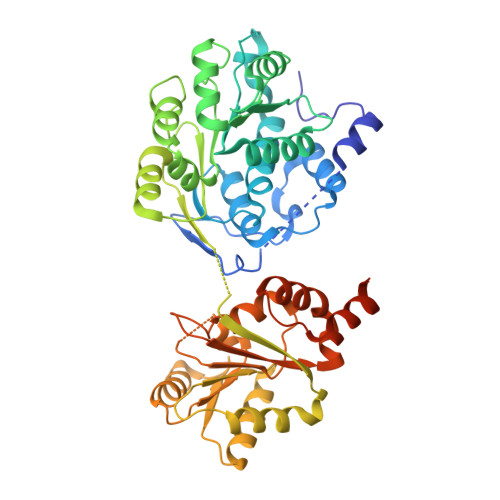
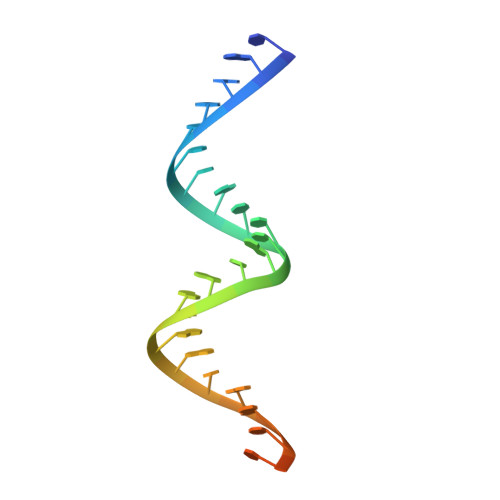
The mechanism of RNA duplex recognition and unwinding by DEAD-box helicase DDX3X.
Song, H., Ji, X.(2019) Nat Commun 10: 3085-3085
- PubMed: 31300642 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-11083-2
- Primary Citation Related Structures:
6O5F - PubMed Abstract:
DEAD-box helicases (DDXs) regulate RNA processing and metabolism by unwinding short double-stranded (ds) RNAs. Sharing a helicase core composed of two RecA-like domains (D1D2), DDXs function in an ATP-dependent, non-processive manner. As an attractive target for cancer and AIDS treatment, DDX3X and its orthologs are extensively studied, yielding a wealth of biochemical and biophysical data, including structures of apo-D1D2 and post-unwound D1D2:single-stranded RNA complex, and the structure of a D2:dsRNA complex that is thought to represent a pre-unwound state. However, the structure of a pre-unwound D1D2:dsRNA complex remains elusive, and thus, the mechanism of DDX action is not fully understood. Here, we describe the structure of a D1D2 core in complex with a 23-base pair dsRNA at pre-unwound state, revealing that two DDXs recognize a 2-turn dsRNA, each DDX mainly recognizes a single RNA strand, and conformational changes induced by ATP binding unwinds the RNA duplex in a cooperative manner.
- Macromolecular Crystallography Laboratory, National Cancer Institute, Frederick, MD, 21702, USA.
Organizational Affiliation: