Discovery and Optimization of Quinolinone Derivatives as Potent, Selective, and Orally Bioavailable Mutant Isocitrate Dehydrogenase 1 (mIDH1) Inhibitors.
Lin, J., Lu, W., Caravella, J.A., Campbell, A.M., Diebold, R.B., Ericsson, A., Fritzen, E., Gustafson, G.R., Lancia Jr., D.R., Shelekhin, T., Wang, Z., Castro, J., Clarke, A., Gotur, D., Josephine, H.R., Katz, M., Diep, H., Kershaw, M., Yao, L., Kauffman, G., Hubbs, S.E., Luke, G.P., Toms, A.V., Wang, L., Bair, K.W., Barr, K.J., Dinsmore, C., Walker, D., Ashwell, S.(2019) J Med Chem 62: 6575-6596
- PubMed: 31199148 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.9b00362
- Primary Citation Related Structures:
6O2Y, 6O2Z - PubMed Abstract:
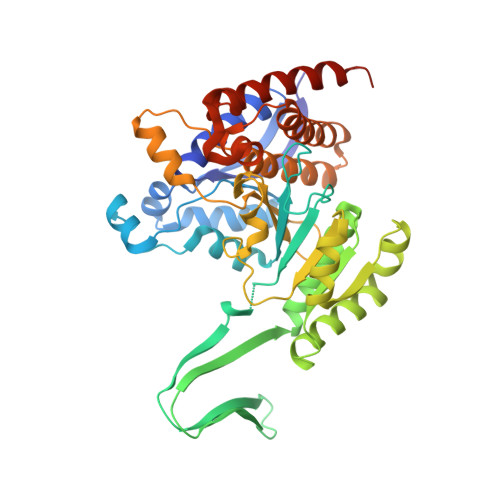
Mutations at the arginine residue (R132) in isocitrate dehydrogenase 1 (IDH1) are frequently identified in various human cancers. Inhibition of mutant IDH1 (mIDH1) with small molecules has been clinically validated as a promising therapeutic treatment for acute myeloid leukemia and multiple solid tumors. Herein, we report the discovery and optimization of a series of quinolinones to provide potent and orally bioavailable mIDH1 inhibitors with selectivity over wild-type IDH1. The X-ray structure of an early lead 24 in complex with mIDH1-R132H shows that the inhibitor unexpectedly binds to an allosteric site. Efforts to improve the in vitro and in vivo absorption, distribution, metabolism, and excretion (ADME) properties of 24 yielded a preclinical candidate 63 . The detailed preclinical ADME and pharmacology studies of 63 support further development of quinolinone-based mIDH1 inhibitors as therapeutic agents in human trials.
- Forma Therapeutics, Inc. , 500 Arsenal Street, Suite 100 , Watertown , Massachusetts 02472 , United States.
Organizational Affiliation: