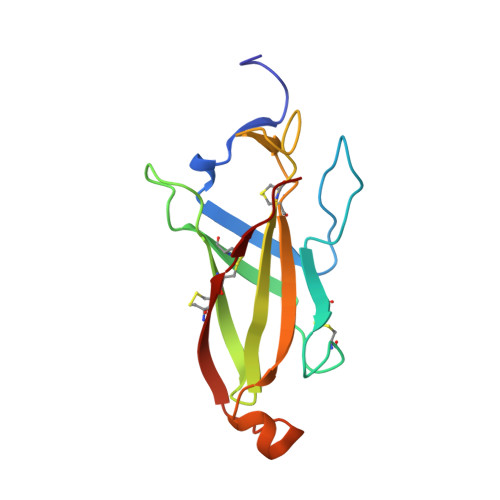
SARAF Luminal Domain Structure Reveals a Novel Domain-Swapped beta-Sandwich Fold Important for SOCE Modulation.
Kimberlin, C.R., Meshcheriakova, A., Palty, R., Raveh, A., Karbat, I., Reuveny, E., Minor Jr., D.L.(2019) J Mol Biology 431: 2869-2883
- PubMed: 31082439 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2019.05.008
- Primary Citation Related Structures:
6O2U, 6O2V, 6O2W - PubMed Abstract:
Store-Operated Calcium Entry (SOCE) plays key roles in cell proliferation, muscle contraction, immune responses, and memory formation. The coordinated interactions of a number of proteins from the plasma and endoplasmic reticulum membranes control SOCE to replenish internal Ca 2+ stores and generate intracellular Ca 2+ signals. SARAF, an endoplasmic reticulum resident component of the SOCE pathway having no homology to any characterized protein, serves as an important brake on SOCE. Here, we describe the X-ray crystal structure of the SARAF luminal domain, SARAF L . This domain forms a novel 10-stranded β-sandwich fold that includes a set of three conserved disulfide bonds, denoted the "SARAF-fold." The structure reveals a domain-swapped dimer in which the last two β-strands (β9 and β10) are exchanged forming a region denoted the "SARAF luminal switch" that is essential for dimerization. Sequence comparisons reveal that the SARAF-fold is highly conserved in vertebrates and in a variety of pathologic fungi. Förster resonance energy transfer experiments using full-length SARAF validate the formation of the domain-swapped dimer in cells and demonstrate that dimerization is reversible. A designed variant lacking the SARAF luminal switch shows that the domain swapping is essential to function and indicates that the SARAF dimer accelerates SOCE inactivation.
- Cardiovascular Research Institute, University of California, San Francisco, CA 94148, USA.
Organizational Affiliation: