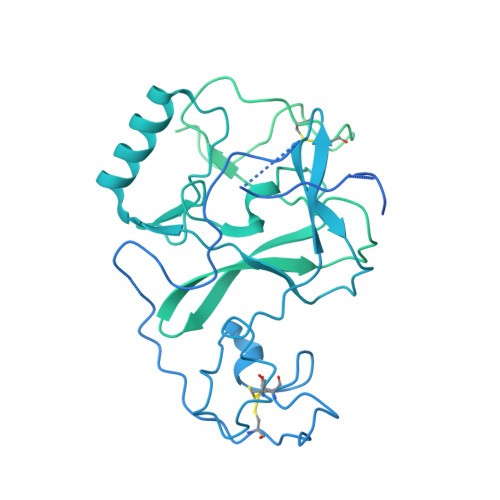
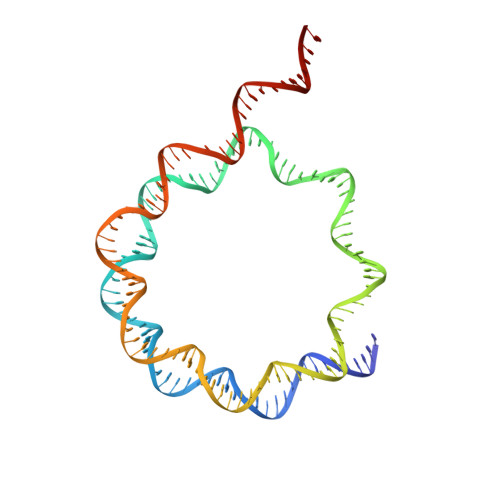
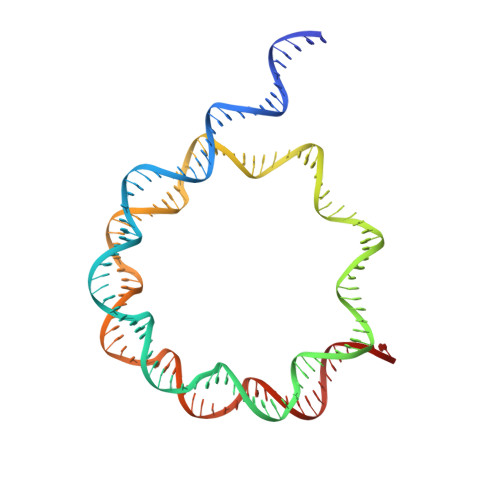
Nucleosome and ubiquitin position Set2 to methylate H3K36.
Bilokapic, S., Halic, M.(2019) Nat Commun 10: 3795-3795
- PubMed: 31439846 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-11726-4
- Primary Citation Related Structures:
6NZO, 6PX1, 6PX3 - PubMed Abstract:
Histone H3 lysine 36 methylation (H3K36me) is a conserved histone modification deposited by the Set2 methyltransferases. Recent findings show that over-expression or mutation of Set2 enzymes promotes cancer progression, however, mechanisms of H3K36me are poorly understood. Set2 enzymes show spurious activity on histones and histone tails, and it is unknown how they obtain specificity to methylate H3K36 on the nucleosome. In this study, we present 3.8 Å cryo-EM structure of Set2 bound to the mimic of H2B ubiquitinated nucleosome. Our structure shows that Set2 makes extensive interactions with the H3 αN, the H3 tail, the H2A C-terminal tail and stabilizes DNA in the unwrapped conformation, which positions Set2 to specifically methylate H3K36. Moreover, we show that ubiquitin contributes to Set2 positioning on the nucleosome and stimulates the methyltransferase activity. Notably, our structure uncovers interfaces that can be targeted by small molecules for development of future cancer therapies.
- Department of Structural Biology, St. Jude Children's Research Hospital, 263 Danny Thomas Place, Memphis, TN, 38105, USA. Silvija.Bilokapic@stjude.org.
Organizational Affiliation: