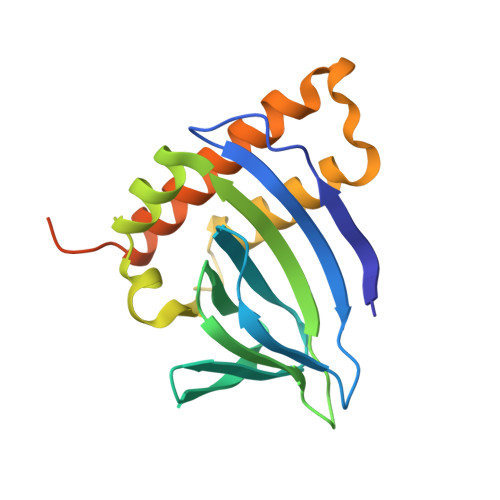
Structures of the transcriptional regulator BgaR, a lactose sensor.
Newman, J., Caron, K., Nebl, T., Peat, T.S.(2019) Acta Crystallogr D Struct Biol 75: 639-646
- PubMed: 31282473 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798319008131
- Primary Citation Related Structures:
6NWH, 6NWJ, 6NWM, 6NWO, 6NX3 - PubMed Abstract:
The structure of BgaR, a transcriptional regulator of the lactose operon in Clostridium perfringens, has been solved by SAD phasing using a mercury derivative. BgaR is an exquisite sensor of lactose, with a binding affinity in the low-micromolar range. This sensor and regulator has been captured bound to lactose and to lactulose as well as in a nominal apo form, and was compared with AraC, another saccharide-binding transcriptional regulator. It is shown that the saccharides bind in the N-terminal region of a jelly-roll fold, but that part of the saccharide is exposed to bulk solvent. This differs from the classical AraC saccharide-binding site, which is mostly sequestered from the bulk solvent. The structures of BgaR bound to lactose and to lactulose highlight how specific and nonspecific interactions lead to a higher binding affinity of BgaR for lactose compared with lactulose. Moreover, solving multiple structures of BgaR in different space groups, both bound to saccharides and unbound, verified that the dimer interface along a C-terminal helix is similar to the dimer interface observed in AraC.
- Biomedical Program, CSIRO, 343 Royal Parade, Parkville, VIC 3052, Australia.
Organizational Affiliation: