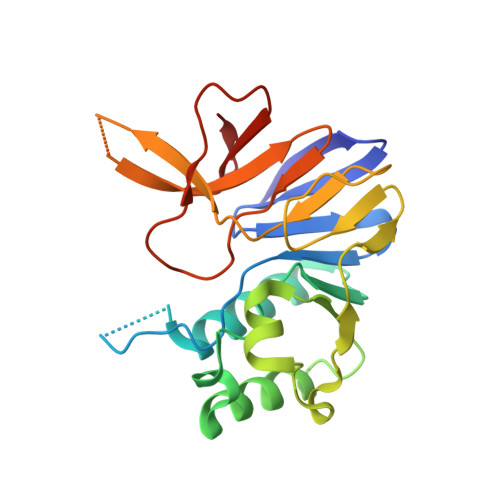
Activity and structure ofPseudomonas putidaMPE, a manganese-dependent single-strand DNA endonuclease encoded in a nucleic acid repair gene cluster.
Ejaz, A., Goldgur, Y., Shuman, S.(2019) J Biological Chem 294: 7931-7941
- PubMed: 30894417 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA119.008049
- Primary Citation Related Structures:
6NVO, 6NVP - PubMed Abstract:
A recently identified and widely prevalent prokaryal gene cluster encodes a suite of enzymes with imputed roles in nucleic acid repair. The enzymes are as follows: MPE, a DNA endonuclease; Lhr-Core, a 3'-5' DNA helicase; LIG, an ATP-dependent DNA ligase; and Exo, a metallo-β-lactamase-family nuclease. Bacterial and archaeal MPE proteins belong to the binuclear metallophosphoesterase superfamily that includes the well-studied DNA repair nucleases Mre11 and SbcD. Here, we report that the Pseudomonas putida MPE protein is a manganese-dependent DNA endonuclease that incises either linear single strands or the single-strand loops of stem-loop DNA structures. MPE has feeble activity on duplex DNA. A crystal structure of MPE at 2.2 Å resolution revealed that the active site includes two octahedrally coordinated manganese ions. Seven signature amino acids of the binuclear metallophosphoesterase superfamily serve as the enzymic metal ligands in MPE: Asp 33 , His 35 , Asp 78 , Asn 112 , His 124 , His 146 , and His 158 A swath of positive surface potential on either side of the active site pocket suggests a binding site for the single-strand DNA substrate. The structure of MPE differs from Mre11 and SbcD in several key respects: (i) MPE is a monomer, whereas Mre11 and SbcD are homodimers; (ii) MPE lacks the capping domain present in Mre11 and SbcD; and (iii) the topology of the β sandwich that comprises the core of the metallophosphoesterase fold differs in MPE vis-à-vis Mre11 and SbcD. We surmise that MPE exemplifies a novel clade of DNA endonuclease within the binuclear metallophosphoesterase superfamily.
- From the Molecular Biology and.
Organizational Affiliation: