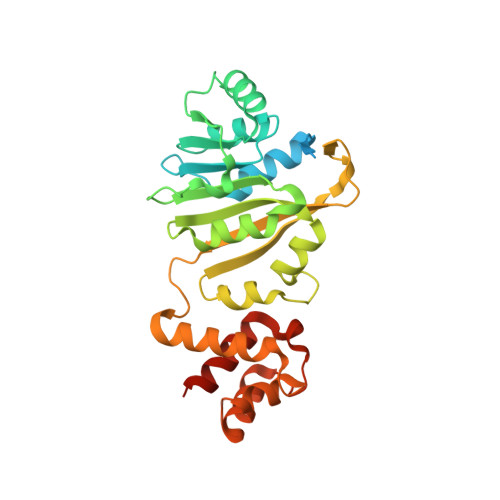
Crystal structure of ErmE - 23S rRNA methyltransferase in macrolide resistance.
Stsiapanava, A., Selmer, M.(2019) Sci Rep 9: 14607-14607
- PubMed: 31601908 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-019-51174-0
- Primary Citation Related Structures:
6NVM - PubMed Abstract:
Pathogens often receive antibiotic resistance genes through horizontal gene transfer from bacteria that produce natural antibiotics. ErmE is a methyltransferase (MTase) from Saccharopolyspora erythraea that dimethylates A2058 in 23S rRNA using S-adenosyl methionine (SAM) as methyl donor, protecting the ribosomes from macrolide binding. To gain insights into the mechanism of macrolide resistance, the crystal structure of ErmE was determined to 1.75 Å resolution. ErmE consists of an N-terminal Rossmann-like α/ß catalytic domain and a C-terminal helical domain. Comparison with ErmC' that despite only 24% sequence identity has the same function, reveals highly similar catalytic domains. Accordingly, superposition with the catalytic domain of ErmC' in complex with SAM suggests that the cofactor binding site is conserved. The two structures mainly differ in the C-terminal domain, which in ErmE contains a longer loop harboring an additional 3 10 helix that interacts with the catalytic domain to stabilize the tertiary structure. Notably, ErmE also differs from ErmC' by having long disordered extensions at its N- and C-termini. A C-terminal disordered region rich in arginine and glycine is also a present in two other MTases, PikR1 and PikR2, which share about 30% sequence identity with ErmE and methylate the same nucleotide in 23S rRNA.
- Department of Cell and Molecular Biology, Uppsala University, BMC, Box 596, SE-751 24, Uppsala, Sweden. alena.stsiapanava@gmail.com.
Organizational Affiliation: