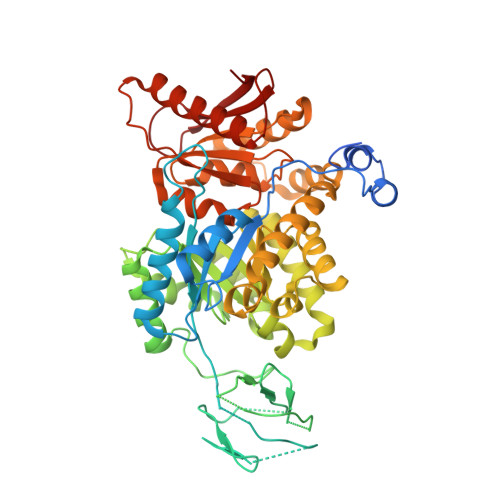
Mechanistic and Structural Insights into Cysteine-Mediated Inhibition of Pyruvate Kinase Muscle Isoform 2.
Srivastava, D., Nandi, S., Dey, M.(2019) Biochemistry 58: 3669-3682
- PubMed: 31386812 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.9b00349
- Primary Citation Related Structures:
6NU1, 6NU5, 6NUB - PubMed Abstract:
Cancer cells regulate key enzymes in the glycolytic pathway to control the glycolytic flux, which is necessary for their growth and proliferation. One of the enzymes is pyruvate kinase muscle isoform 2 (PKM2), which is allosterically regulated by various small molecules. Using detailed biochemical and kinetic studies, we demonstrate that cysteine inhibits wild-type (wt) PKM2 by shifting from an active tetramer to a mixture of a tetramer and a less active dimer/monomer equilibrium and that the inhibition is dependent on cysteine concentration. The cysteine-mediated PKM2 inhibition is reversed by fructose 1,6-bisphosphate, an allosteric activator of PKM2. Furthermore, kinetic studies using two dimeric PKM2 variants, S437Y PKM2 and G415R PKM2, show that the reversal is caused by the tetramerization of wtPKM2. The crystal structure of the wtPKM2-Cys complex was determined at 2.25 Å, which showed that cysteine is held to the amino acid binding site via its main chain groups, similar to that observed for phenylalanine, alanine, serine, and tryptophan. Notably, ligand binding studies using fluorescence and isothermal titration calorimetry show that the presence of phosphoenolpyruvate alters the binding affinities of amino acids for wtPKM2 and vice versa, thereby unravelling the existence of a functionally bidirectional coupling between the amino acid binding site and the active site of wtPKM2.
- Department of Chemistry , The University of Iowa , Iowa City , Iowa 52242 , United States.
Organizational Affiliation: