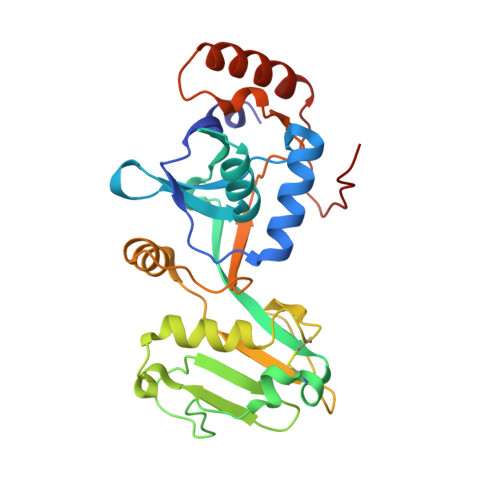
The heme-sensitive regulator SbnI has a bifunctional role in staphyloferrin B production by Staphylococcus aureus .
Verstraete, M.M., Morales, L.D., Kobylarz, M.J., Loutet, S.A., Laakso, H.A., Pinter, T.B., Stillman, M.J., Heinrichs, D.E., Murphy, M.E.P.(2019) J Biological Chem 294: 11622-11636
- PubMed: 31197035 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA119.007757
- Primary Citation Related Structures:
5UJD, 6NR6 - PubMed Abstract:
Staphylococcus aureus infection relies on iron acquisition from its host. S. aureus takes up iron through heme uptake by the iron-responsive surface determinant (Isd) system and by the production of iron-scavenging siderophores. Staphyloferrin B (SB) is a siderophore produced by the 9-gene sbn gene cluster for SB biosynthesis and efflux. Recently, the ninth gene product, SbnI, was determined to be a free l-serine kinase that produces O- phospho-l-serine (OPS), a substrate for SB biosynthesis. Previous studies have also characterized SbnI as a DNA-binding regulatory protein that senses heme to control sbn gene expression for SB synthesis. Here, we present crystal structures at 1.9-2.1 Å resolution of a SbnI homolog from Staphylococcus pseudintermedius (SpSbnI) in both apo form and in complex with ADP, a product of the kinase reaction; the latter confirmed the active-site location. The structures revealed that SpSbnI forms a dimer through C-terminal domain swapping and a dimer of dimers through intermolecular disulfide formation. Heme binding had only a modest effect on SbnI enzymatic activity, suggesting that its two functions are independent and structurally distinct. We identified a heme-binding site and observed catalytic heme transfer between a heme-degrading protein of the Isd system, IsdI, and SbnI. These findings support the notion that SbnI has a bifunctional role contributing precursor OPS to SB synthesis and directly sensing heme to control expression of the sbn locus. We propose that heme transfer from IsdI to SbnI enables S. aureus to control iron source preference according to the sources available in the environment.
- Department of Microbiology and Immunology, University of British Columbia, Vancouver, BC V6T 1Z3, Canada.
Organizational Affiliation: