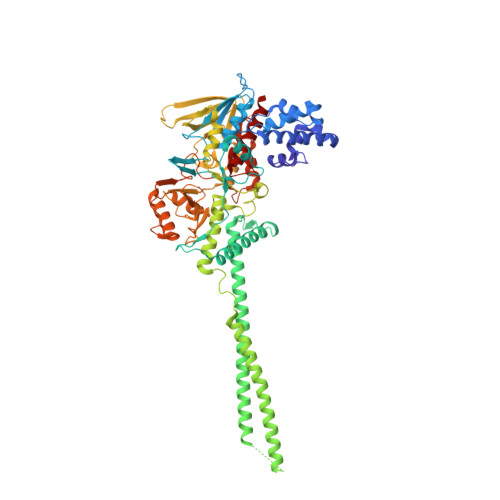
Lysine-Specific Histone Demethylase 1A Regulates Macrophage Polarization and Checkpoint Molecules in the Tumor Microenvironment of Triple-Negative Breast Cancer.
Tan, A.H.Y., Tu, W., McCuaig, R., Hardy, K., Donovan, T., Tsimbalyuk, S., Forwood, J.K., Rao, S.(2019) Front Immunol 10: 1351-1351
- PubMed: 31249575 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/fimmu.2019.01351
- Primary Citation Related Structures:
6NQM, 6NQU, 6NR5 - PubMed Abstract:
Macrophages play an important role in regulating the tumor microenvironment (TME). Here we show that classical (M1) macrophage polarization reduced expression of LSD1, nuclear REST corepressor 1 (CoREST), and the zinc finger protein SNAIL. The LSD1 inhibitor phenelzine targeted both the flavin adenine dinucleotide (FAD) and CoREST binding domains of LSD1, unlike the LSD1 inhibitor GSK2879552, which only targeted the FAD domain. Phenelzine treatment reduced nuclear demethylase activity and increased transcription and expression of M1-like signatures both in vitro and in a murine triple-negative breast cancer model. Overall, the LSD1 inhibitors phenelzine and GSK2879552 are useful tools for dissecting the contribution of LSD1 demethylase activity and the nuclear LSD1-CoREST complex to switching macrophage polarization programs. These findings suggest that inhibitors must have dual FAD and CoREST targeting abilities to successfully initiate or prime macrophages toward an anti-tumor M1-like phenotype in triple-negative breast cancer.
- Epigenetics and Transcription Laboratory Melanie Swan Memorial Translational Centre, Sci-Tech, University of Canberra, Canberra, ACT, Australia.
Organizational Affiliation: