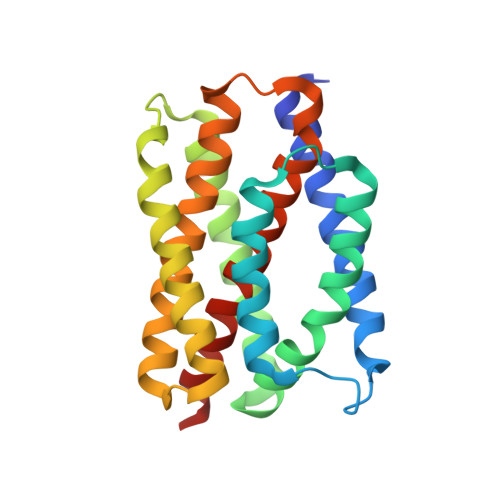
Ion and pH Sensitivity of a TMBIM Ca2+Channel.
Guo, G., Xu, M., Chang, Y., Luyten, T., Seitaj, B., Liu, W., Zhu, P., Bultynck, G., Shi, L., Quick, M., Liu, Q.(2019) Structure 27: 1013-1021.e3
- PubMed: 30930064 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2019.03.003
- Primary Citation Related Structures:
6NQ7, 6NQ8, 6NQ9 - PubMed Abstract:
The anti-apoptotic transmembrane Bax inhibitor motif (TMBIM) containing protein family regulates Ca 2+ homeostasis, cell death, and the progression of diseases including cancers. The recent crystal structures of the TMBIM homolog BsYetJ reveal a conserved Asp171-Asp195 dyad that is proposed in regulating a pH-dependent Ca 2+ translocation. Here we show that BsYetJ mediates Ca 2+ fluxes in permeabilized mammalian cells, and its interaction with Ca 2+ is sensitive to protons and other cations. We report crystal structures of BsYetJ in additional states, revealing the flexibility of the dyad in a closed state and a pore-opening mechanism. Functional studies show that the dyad is responsible for both Ca 2+ affinity and pH dependence. Computational simulations suggest that protonation of Asp171 weakens its interaction with Arg60, leading to an open state. Our integrated analysis provides insights into the regulation of the BsYetJ Ca 2+ channel that may inform understanding of human TMBIM proteins regarding their roles in cell death and diseases.
- Biology Department, Brookhaven National Laboratory, Upton, NY 11973, USA; NSLS-II, Brookhaven National Laboratory, Upton, NY 11973, USA.
Organizational Affiliation: