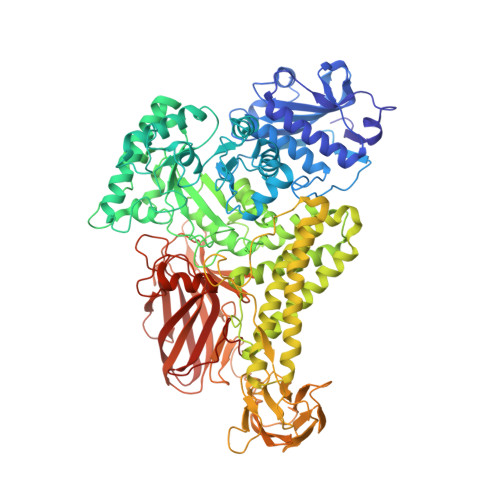
Structural characterization of the family GH115 alpha-glucuronidase from Amphibacillus xylanus yields insight into its coordinated action with alpha-arabinofuranosidases.
Yan, R., Wang, W., Vuong, T.V., Xiu, Y., Skarina, T., Di Leo, R., Gatenholm, P., Toriz, G., Tenkanen, M., Stogios, P.J., Master, E.R.(2021) N Biotechnol
- PubMed: 33486119
- DOI: https://doi.org/10.1016/j.nbt.2021.01.005
- Primary Citation Related Structures:
6NPS - PubMed Abstract:
The coordinated action of carbohydrate-active enzymes has mainly been evaluated for the purpose of complete saccharification of plant biomass (lignocellulose) to sugars. By contrast, the coordinated action of accessory hemicellulases on xylan debranching and recovery is less well characterized. Here, the activity of two family GH115 α-glucuronidases (SdeAgu115A from Saccharophagus degradans, and AxyAgu115A from Amphibacillus xylanus) on spruce arabinoglucuronoxylan (AGX) was evaluated in combination with an α-arabinofuranosidase from families GH51 (AniAbf51A, aka E-AFASE from Aspergillus niger) and GH62 (SthAbf62A from Streptomyces thermoviolaceus). The α-arabinofuranosidases boosted (methyl)-glucuronic acid release by SdeAgu115A by approximately 50 % and 30 %, respectively. The impact of the α-arabinofuranosidases on AxyAgu115A activity was comparatively low, motivating its structural characterization. The crystal structure of AxyAgu115A revealed increased length and flexibility of the active site loop compared to SdeAgu115A. This structural difference could explain the ability of AxyAgu115A to accommodate more highly substituted arabinoglucuronoxylan, and inform enzyme selections for improved AGX recovery and use.
- Department of Chemical Engineering and Applied Chemistry, University of Toronto, 200 College Street, Toronto, Ontario, M5S 3E5, Canada.
Organizational Affiliation: