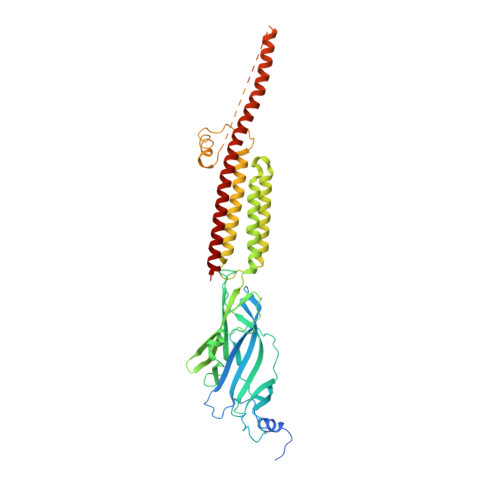
Molecular mechanism of setron-mediated inhibition of full-length 5-HT3Areceptor.
Basak, S., Gicheru, Y., Kapoor, A., Mayer, M.L., Filizola, M., Chakrapani, S.(2019) Nat Commun 10: 3225-3225
- PubMed: 31324772 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-11142-8
- Primary Citation Related Structures:
6NP0 - PubMed Abstract:
Serotonin receptor (5-HT 3A R) is the most common therapeutic target to manage the nausea and vomiting during cancer therapies and in the treatment of irritable bowel syndrome. Setrons, a class of competitive antagonists, cause functional inhibition of 5-HT 3A R in the gastrointestinal tract and brainstem, acting as effective anti-emetic agents. Despite their prevalent use, the molecular mechanisms underlying setron binding and inhibition of 5-HT 3A R are not fully understood. Here, we present the structure of granisetron-bound full-length 5-HT 3A R solved by single-particle cryo-electron microscopy to 2.92 Å resolution. The reconstruction reveals the orientation of granisetron in the orthosteric site with unambiguous density for interacting sidechains. Molecular dynamics simulations and electrophysiology confirm the granisetron binding orientation and the residues central for ligand recognition. Comparison of granisetron-bound 5-HT 3A R with the apo and serotonin-bound structures, reveals key insights into the mechanism underlying 5-HT 3A R inhibition.
- Department of Physiology and Biophysics, Case Western Reserve University, Cleveland, OH, 44106-4970, USA.
Organizational Affiliation: