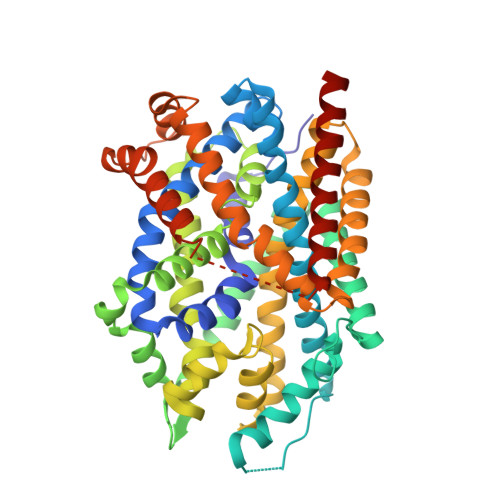
Structural, functional, and behavioral insights of dopamine dysfunction revealed by a deletion inSLC6A3.
Campbell, N.G., Shekar, A., Aguilar, J.I., Peng, D., Navratna, V., Yang, D., Morley, A.N., Duran, A.M., Galli, G., O'Grady, B., Ramachandran, R., Sutcliffe, J.S., Sitte, H.H., Erreger, K., Meiler, J., Stockner, T., Bellan, L.M., Matthies, H.J.G., Gouaux, E., Mchaourab, H.S., Galli, A.(2019) Proc Natl Acad Sci U S A 116: 3853-3862
- PubMed: 30755521 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1816247116
- Primary Citation Related Structures:
6NLE - PubMed Abstract:
The human dopamine (DA) transporter (hDAT) mediates clearance of DA. Genetic variants in hDAT have been associated with DA dysfunction, a complication associated with several brain disorders, including autism spectrum disorder (ASD). Here, we investigated the structural and behavioral bases of an ASD-associated in-frame deletion in hDAT at N336 (∆N336). We uncovered that the deletion promoted a previously unobserved conformation of the intracellular gate of the transporter, likely representing the rate-limiting step of the transport process. It is defined by a "half-open and inward-facing" state (HOIF) of the intracellular gate that is stabilized by a network of interactions conserved phylogenetically, as we demonstrated in hDAT by Rosetta molecular modeling and fine-grained simulations, as well as in its bacterial homolog leucine transporter by electron paramagnetic resonance analysis and X-ray crystallography. The stabilization of the HOIF state is associated both with DA dysfunctions demonstrated in isolated brains of Drosophila melanogaster expressing hDAT ∆N336 and with abnormal behaviors observed at high-time resolution. These flies display increased fear, impaired social interactions, and locomotion traits we associate with DA dysfunction and the HOIF state. Together, our results describe how a genetic variation causes DA dysfunction and abnormal behaviors by stabilizing a HOIF state of the transporter.
- Department of Molecular Physiology & Biophysics, Vanderbilt University, Nashville, TN 37232.
Organizational Affiliation: