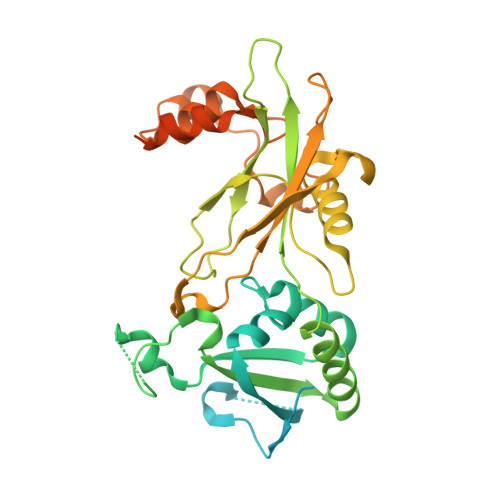
Structures of MERS1, the 5' processing enzyme of mitochondrial mRNAs inTrypanosoma brucei.
Schumacher, M.A., Henderson, M., Zeng, W.(2020) RNA 26: 69-82
- PubMed: 31704716 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1261/rna.072231.119
- Primary Citation Related Structures:
6NL1, 6P5R, 6U9X - PubMed Abstract:
Most mitochondrial mRNAs are transcribed as polycistronic precursors that are cleaved by endonucleases to produce mature mRNA transcripts. However, recent studies have shown that mitochondrial transcripts in the kinetoplastid protozoan, Trypanosoma brucei , are transcribed individually. Also unlike most mitochondrial mRNAs, the 5' end of these transcripts harbor a triphosphate that is hydrolyzed. This modification is carried out by a putative Nudix hydrolase called MERS1. The Nudix motif in MERS1 is degenerate, lacking a conserved glutamic acid, thus it is unclear how it may bind its substrates and whether it contains a Nudix fold. To obtain insight into this unusual hydrolase, we determined structures of apo, GTP-bound and RNA-bound T. brucei MERS1 to 2.30 Å, 2.45 Å, and 2.60 Å, respectively. The MERS1 structure has a unique fold that indeed contains a Nudix motif. The nucleotide bound structures combined with binding studies reveal that MERS1 shows preference for RNA sequences with a central guanine repeat which it binds in a single-stranded conformation. The apo MERS1 structure indicates that a significant portion of its nucleotide binding site folds upon substrate binding. Finally, a potential interaction region for a binding partner, MERS2, that activates MERS1 was identified. The MERS2-like peptide inserts a glutamate near the missing Nudix acidic residue in the RNA binding pocket, suggesting how the enzyme may be activated. Thus, the combined studies reveal insight into the structure and enzyme properties of MERS1 and its substrate-binding activities.
- Department of Biochemistry, Duke University School of Medicine, Durham, North Carolina 27710, USA.
Organizational Affiliation: