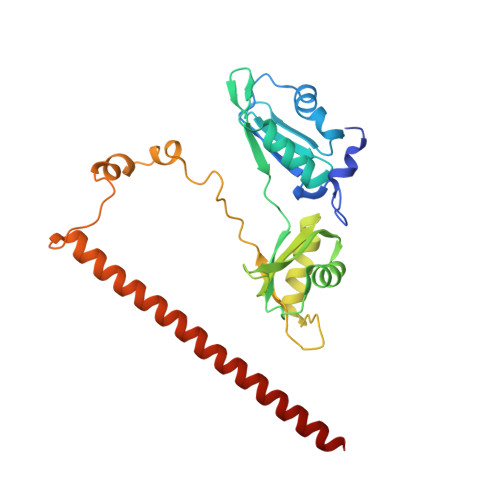
A new crystal structure and small-angle X-ray scattering analysis of the homodimer of human SFPQ.
Hewage, T.W., Caria, S., Lee, M.(2019) Acta Crystallogr F Struct Biol Commun 75: 439-449
- PubMed: 31204691 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X19006599
- Primary Citation Related Structures:
6NCQ - PubMed Abstract:
Splicing factor proline/glutamine-rich (SFPQ) is an essential RNA-binding protein that is implicated in many aspects of nuclear function. The structures of SFPQ and two paralogs, non-POU domain-containing octamer-binding protein and paraspeckle component 1, from the Drosophila behavior human splicing protein family have previously been characterized. The unusual arrangement of the four domains, two RNA-recognition motifs (RRMs), a conserved region termed the NonA/paraspeckle (NOPS) domain and a C-terminal coiled coil, in the intertwined dimer provides a potentially unique RNA-binding surface. However, the molecular details of how the four RRMs in the dimeric SFPQ interact with RNA remain to be characterized. Here, a new crystal structure of the dimerization domain of human SFPQ in the C-centered orthorhombic space group C222 1 with one monomer in the asymmetric unit is presented. Comparison of the new crystal structure with the previously reported structure of SFPQ and analysis of the solution small-angle X-scattering data revealed subtle domain movements in the dimerization domain of SFPQ, supporting the concept of multiple conformations of SFPQ in equilibrium in solution. The domain movement of RRM1, in particular, may reflect the complexity of the RNA substrates of SFPQ. Taken together, the crystal and solution structure analyses provide a molecular basis for further investigation into the plasticity of nucleic acid binding by SFPQ in the absence of the structure in complex with its cognate RNA-binding partners.
- Department of Biochemistry and Genetics, La Trobe Institute for Molecular Science, La Trobe University, Melbourne, Victoria 3086, Australia.
Organizational Affiliation: